Versions Compared
Key
- This line was added.
- This line was removed.
- Formatting was changed.
Overview
BioNetDB models biology data as a network of nodes and relations.Biology data comes from different formats and sources it comprises system biology data from Reactome, annotation data from CellBase and human genetic variations from healthcare centers' clinical data. BioNetDB relies on Neo4j graph database that allows users to access biological data using the Cypher query language (similar to SQL in relational databases). Neo4j is highly optimized for queries and it is scalable and reliable.
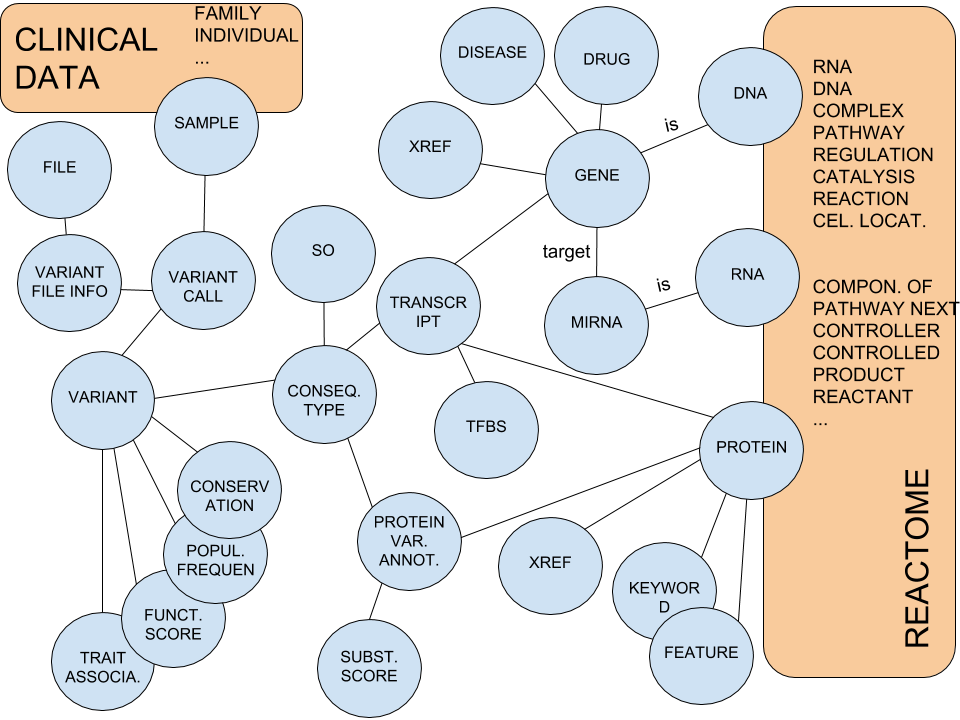
Modelling
ThGenes
FILE
VARIANT_CALL
VARIANT_FILE_INFO
DRUG
DISEASE
GENE
MIRNA
DNA
VARIANT_FILE_INFO
CATALYSISCELLULAR_LOCATION
COMPLEX
CONSEQUENCE_TYPE
CONSERVATION
FUNCTIONAL_SCORE
PATHWAY
POPULATION_FREQUENCY
PROTEIN
PROTEIN_FEATURE
PROTEIN_KEYWORD
PROTEIN_VARIANT_ANNOTATION
REACTION
REGULATION
RNA
SAMPLE
SMALL_MOLECULE
SO
SUBSTITUTION_SCORE
TFBS
TRAIT_ASSOCIATION
TRANSCRIPT
UNDEFINED
VARIANT
XREF
Genes
TranscriptsTranscripts
Transcript node properties are:
id
name
biotype
chromosome
- start
- end
- strand
proteinId
genomicCodingEnd
genomicCodingStart
annotationFlags
cdnaCodingEnd
cdnaCodingStart
cdsLength
description
status
Image below shows the transcript relationships:
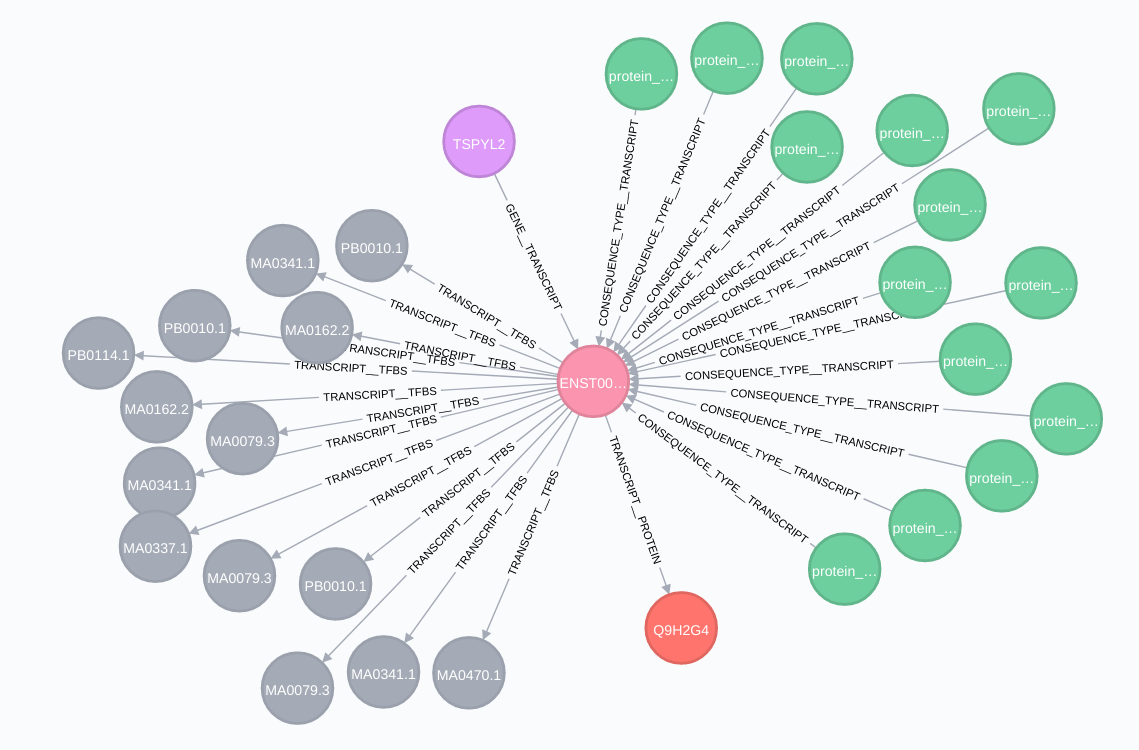
Proteins
Protein Complex
Variants
Regulation
Pathway
Table of Contents:
| Table of Contents | ||
|---|---|---|
|