BioNetDB models biology data as a network of nodes and relations.Biology data comes from different formats and sources it comprises system biology data from Reactome, annotation data from CellBase and human genetic variations from healthcare centers' clinical data. BioNetDB relies on Neo4j graph database that allows users to access biological data using the Cypher query language (similar to SQL in relational databases).
The figure below shows BioNetDB nodes with their labels. for clarity, some labels have been shortened:
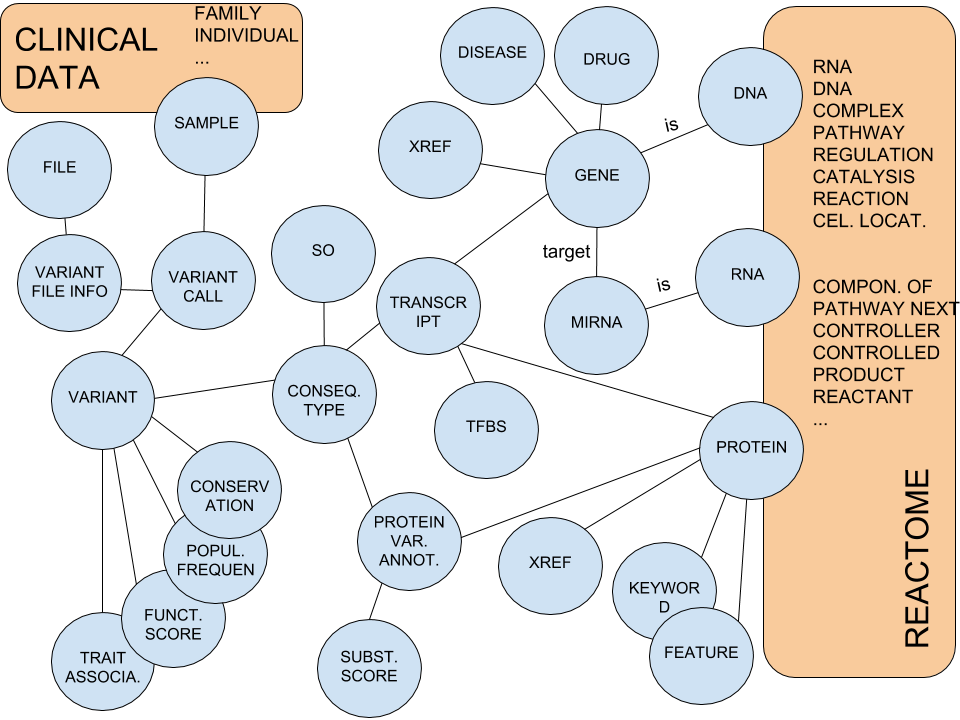
Shortened labes in the previous figure:
| Shortened label | Node label |
|---|---|
1 POPUL. FREQUEN | POPULATION_FREQUENCY |
| 2 FUNCT. SCORE | FUNCTIONAL_SCORE |
3 TRAIT ASSOCIA. | TRAIT_ASSOCIATION |
4 CONSEQ. TYPE | CONSEQUENCE_TYPE |
| 5 PROTEIN VAR. ANNOT. | PROTEIN_VARIANT_ANNOTATION |
| 6 SUBST. SCORE | SUBSTITUTION_SCORE |
| 7 KEYWORD | PROTEIN_KEYWORD |
| 8 FEATURE | PROTEIN_FEATURE |
Modelling
This section lists the main nodes of the BioNetDB network data model and for each of them, its properties and relationships are shown.
Genes
Gene node properties:
uid
id
name
chromosome
start
end
strand
- biotype
description:
source
status
Gene relationships:
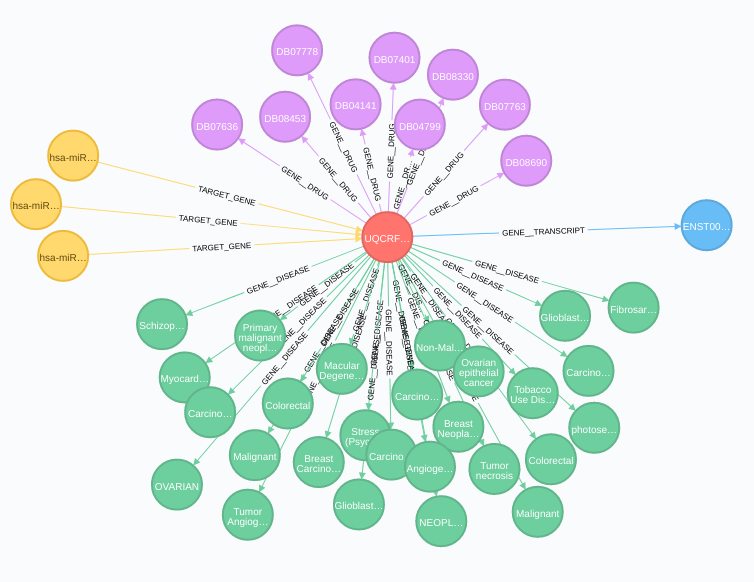
Transcripts
Transcript node properties:
uid
id
name
biotype
chromosome
- start
- end
- strand
proteinId
genomicCodingEnd
genomicCodingStart
annotationFlags
cdnaCodingEnd
cdnaCodingStart
cdsLength
description
status
Transcript relationships (transcript node in pink):
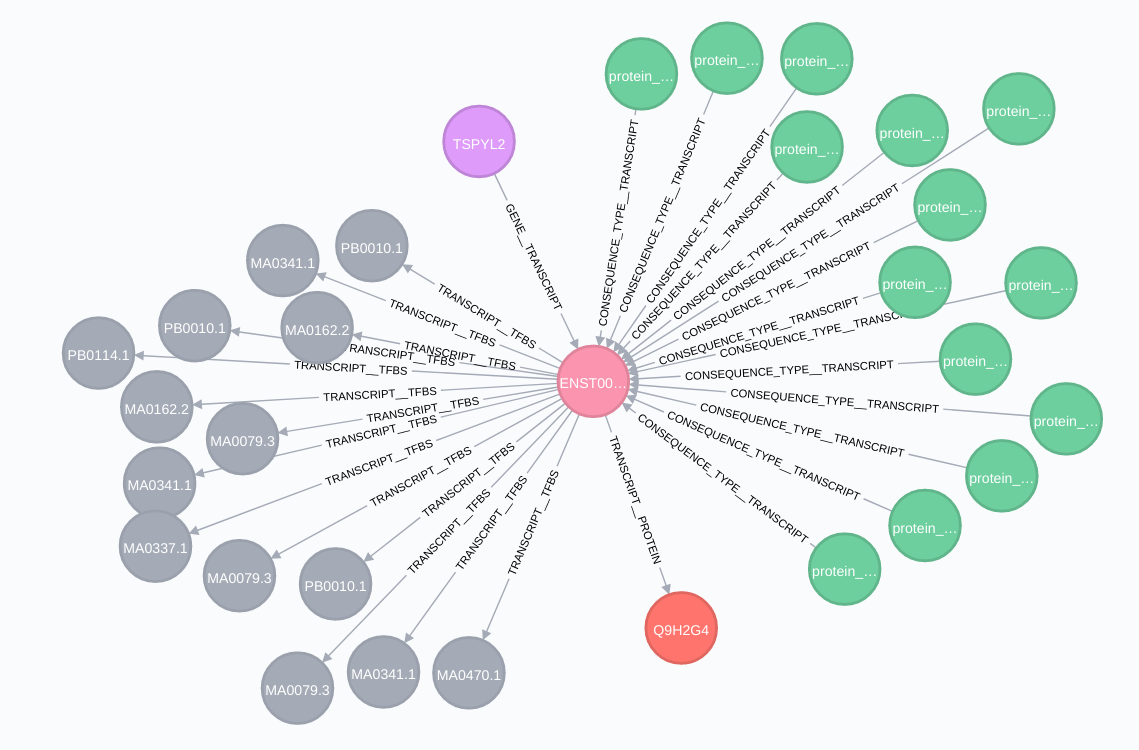
Proteins
Protein node properties:
uid
id
name
accession
dataset
- evidence
- proteinExistence
Protein relationships:
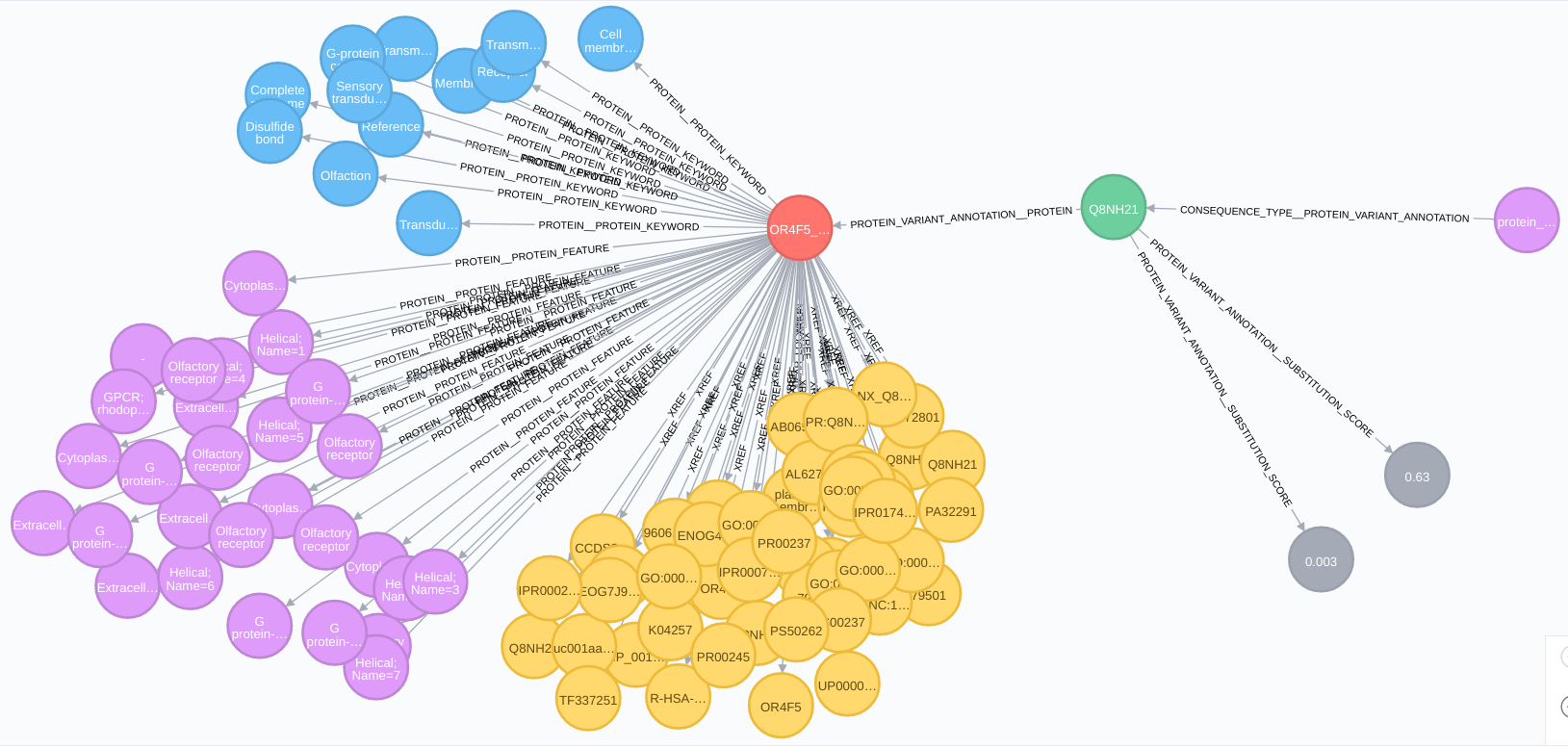
Protein complex
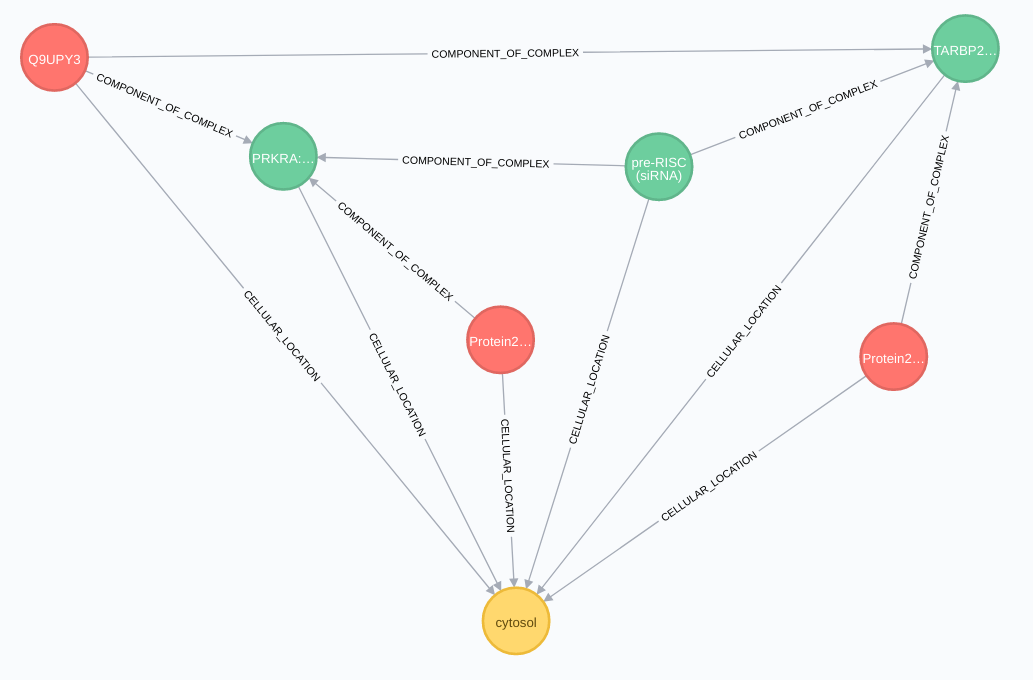
Variants
Variant node properties:
uid
id
- name
- chromosome
- start
- end
- strand
- type
- alternate
- reference
alternativeNames
Variant relationships:
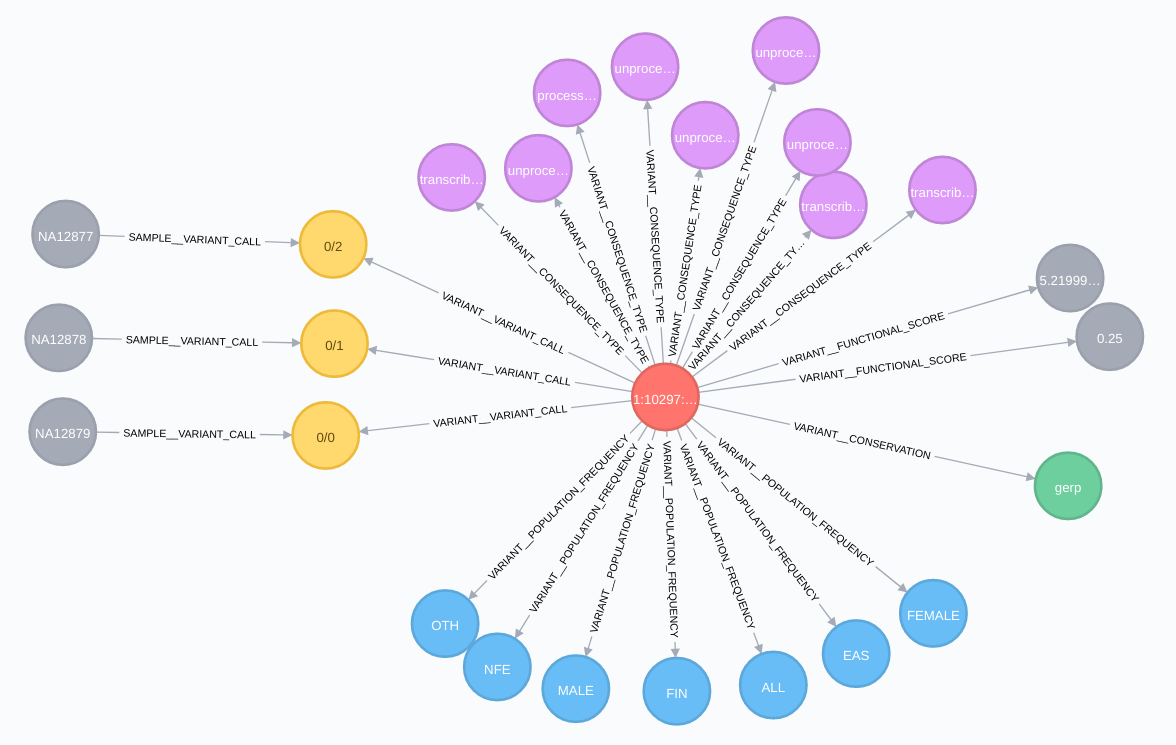
Regulation
Regulation node properties:
uid
id
- name
Regulation relationships:
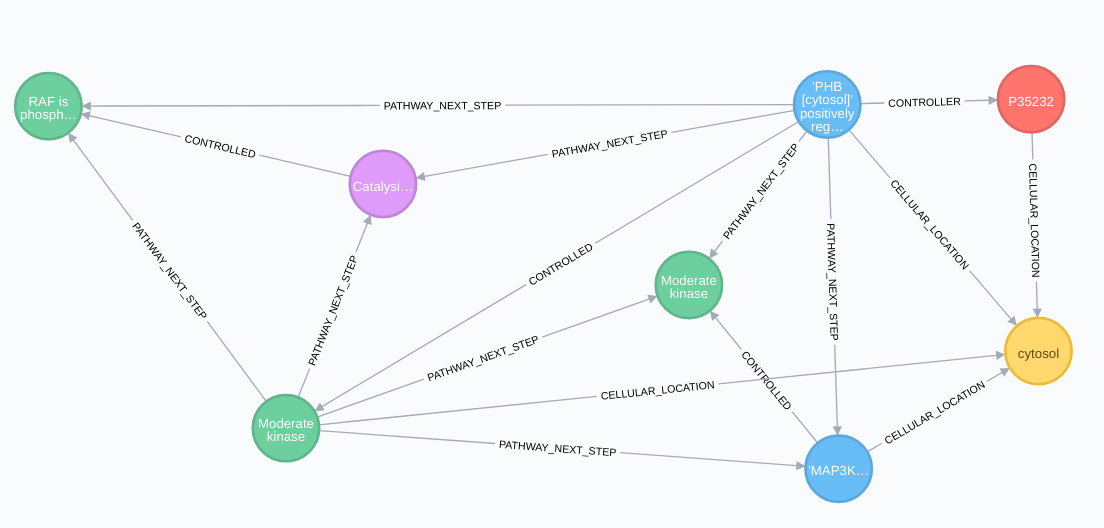
Pathway
Pathway node properties:
uid
id
- name
Pathway relationships (pathway nodes in yellow):
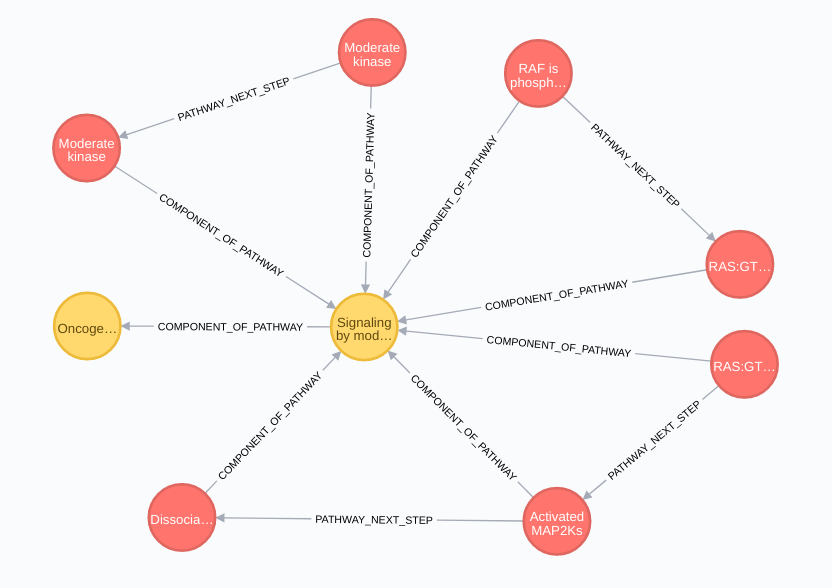
Table of Contents: