Client-Server Model
HGVA is implemented following a Client-Server architecture. The server uses OpenCB OpenCGA project to load and index variant datasets (VCF and gVCF files), OpenCGA server provides a complete REST API to query metadata and variants. The client side implements three different user interfaces. First, a rich web-based data mining application based on OpenCB IVA project. Second, three client libraries for Java, Python and Javascript. Third, a command-line interface. Client libraries and command-line can query both metadata and variants and are part of OpenCGA project.

OpenCGA Server
OpenCGA is an open-source project that aims to provide a Big Data storage engine and analysis framework for genomic scale data analysis of hundreds of terabytes. It implements different components:
- Catalog to store metadata
- Variant storage engine to provide real-time queries to big data in genomics. This can use MongoDB or HBase together with Solr. A Redis server is also used to cache queries.
- A complete RESTful API and gRPC for variant and alignment (BAM) streaming
- Client libraries and command-line to query data
HGVA uses a small cluster for OpenCGA installation. This consist of three servers for MongoDB and a single Solr server. HAProxy is used to balance queries to two Tomcat servers.
Client User Interfaces
Client user interfaces to HGVA include a rich web application based on IVA, client libraries in Java, Python and JavaScript, and a Command Line Interface. All of these make intensive use of HGVA's RESTful web services (taken from OpenCGA), which are accessible through an HAProxy load balancer.
OpenCB IVA
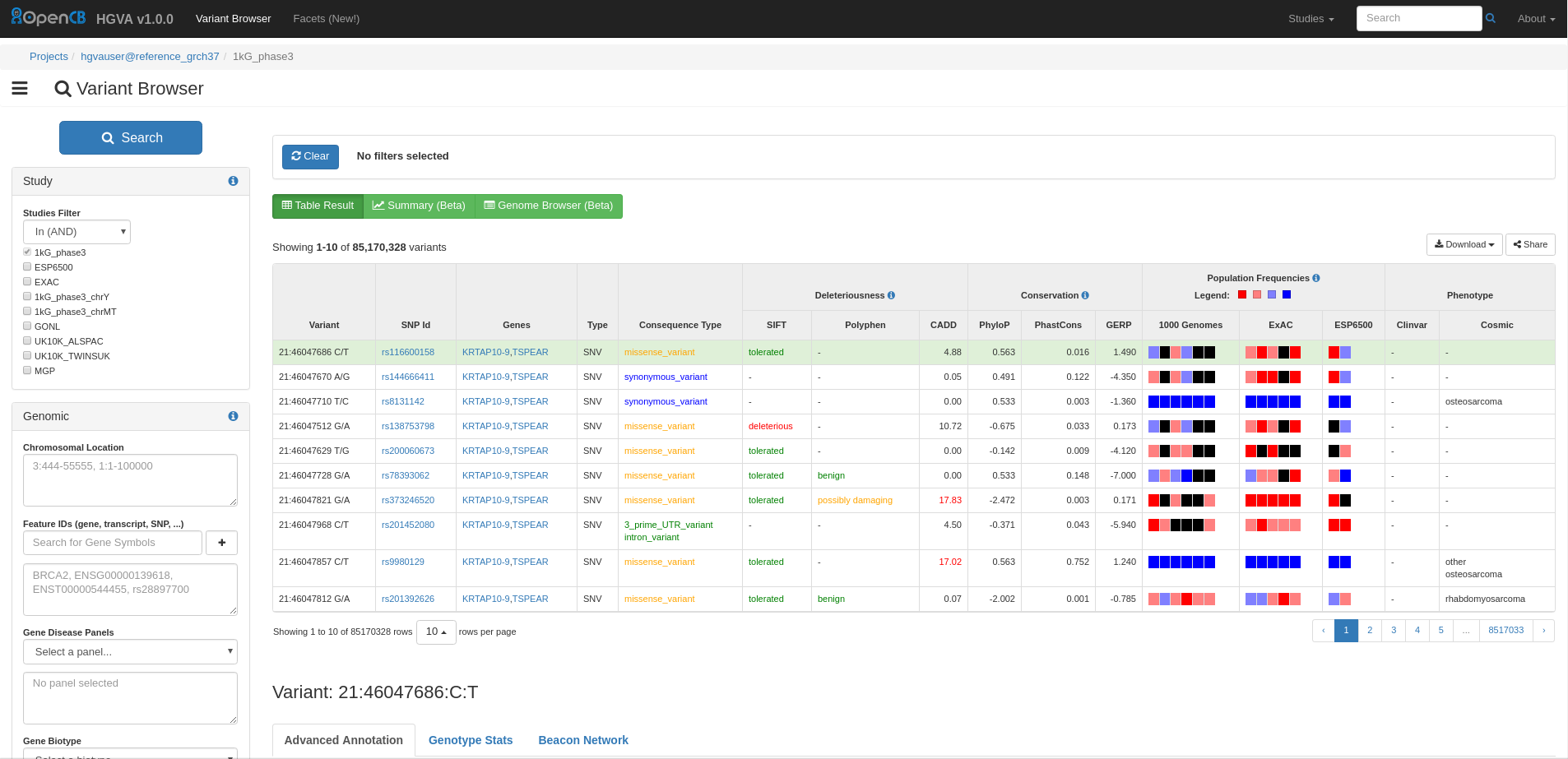
IVA is a highly customisable web application for Interactive Variant Analysis (IVA). It consists of several tools, HGVA activates two of them: Variant Browser and Facets. You can execute complex queries in IVA using any variant annotation including full-text search for disease descriptions.
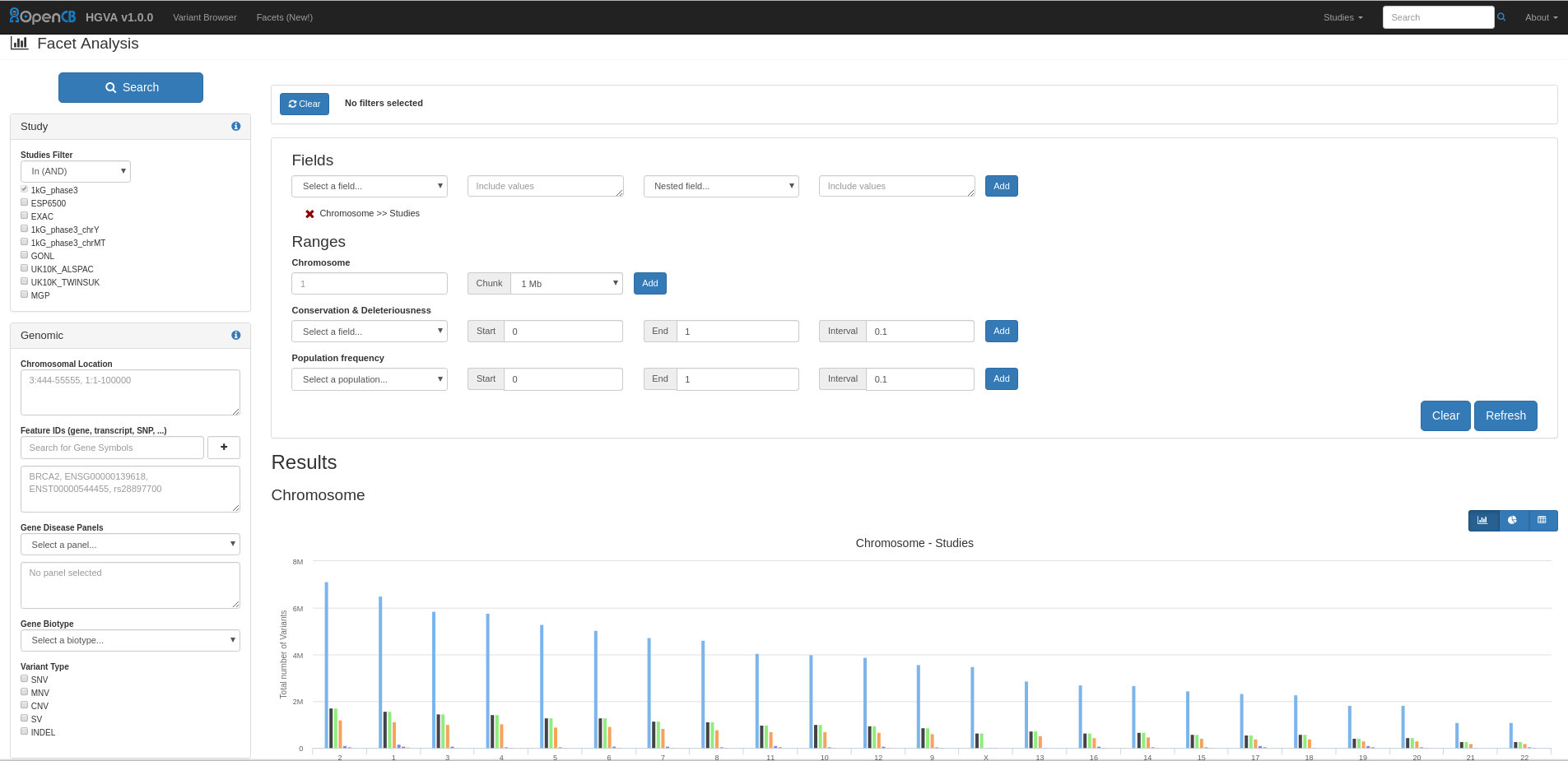
With Facets you can perform different aggregations of data: