Index pipeline is the process of ingesting data into an OpenCGA-Storage backend. We define a general pipeline that is used and extended for the supported bioformats like variants and alignments. This pipeline can be extended by additional steps of enrichment, which will be highly dependent on the file format. At the end, the data may be filtered to be visualized, or used as analysis input data.
This concept is represented in Catalog to help the tracking of this status in different files.
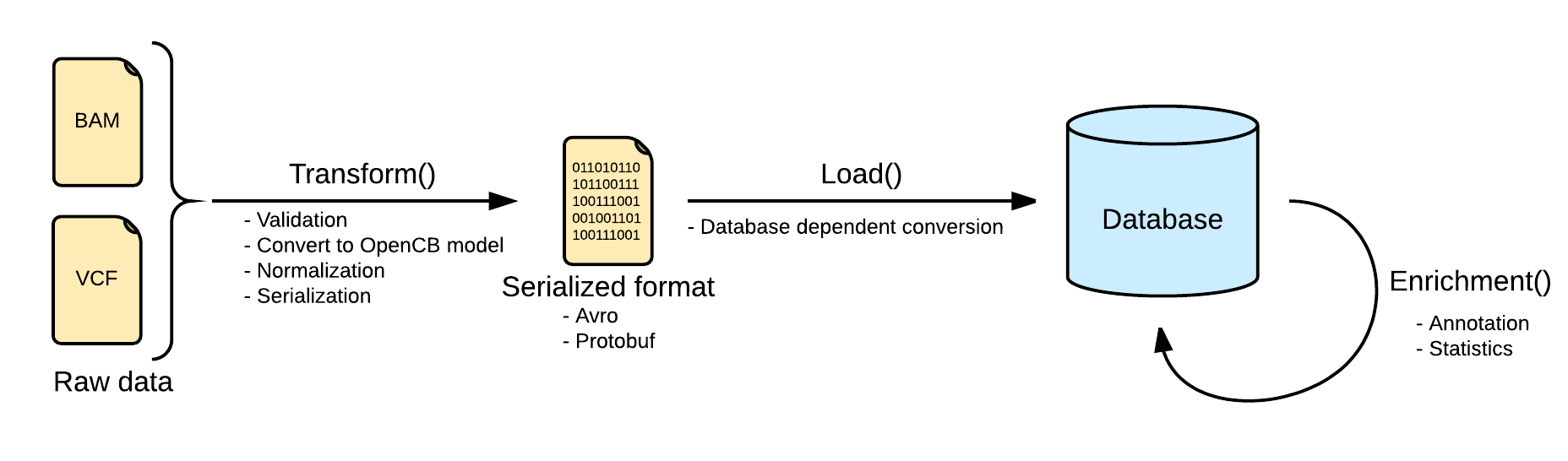
Indexing data pipeline consists in three steps, transforming the input raw data into an intermediate format, loading it into the selected database, depending on the implementation, and adding more information to the loaded data by calculating statistics or adding extra information like annotation.
Transform
The first and one of the most important steps is the transformation. At this point, the pipeline ensures that the input file is valid and can be loaded into the database. The input file is read and converted into the OpenCB models, defined in Biodata. One single input file may generate more than one file, separating the data from the metadata. See Data Models for more info.
Depending on the input data the process will be more or less complex, and, at the end, the file will be serialized into disk using some serialization schema like Avro, Protobuf or Json in some cases.
As the transformation stage grants that the data is valid and can be loaded, the load stage can not start until the transformation has finished.
This step is shared between all the storage engine implementations of the same bioformat, so the result should be valid for any implementation.
Load
This step is intended to be as fast as possible, to avoid unnecessary downtimes in the database due to the work load. Because of this, all the convert and validate operations are made in the previous step.
Most of the storage engines are not going to load directly the opencb models, and some more engine dependent transformations are still expected. The storage engines grant that any valid instance of the input data model can be transformed and loaded.
In some scenarios the load step may be done in two steps, loading first into an intermediate system a batch of files, and then, move all of them to the real storage system. This could improve the loading speed by consuming more storage resources.
Enrichment
Despite input file formats contains a lot of interesting information, some of it is not directly available, and has to be calculated or fetched from external services.
Most of the storage engines provide mechanisms to calculate some statistics (either per record or aggregated) and store them back in the database to help the filtering process. By doing this, we can speed up some queries against pre-calculated statistics.
Some other information can not be inferred from the input data, and has to be fetched from external annotation services like Cellbase