Overview
Understanding the URL
The general format of the REST API web services is:
http://HOST_URL/webservices/rest/{apiVersion}/{resource}/{id(s)}/{endpoint}?{options} |
where HOST_URL is the name of the host machine and the name of Java war file deployed in web server (eg. tomcat) over that server for example http://bioinfo.hpc.cam.ac.uk/opencga-demo/
Entities inside the curly braces { } are the web service parameters, and they are treated as variables. For example the following URL:
http://HOST_URL/webservices/rest/v1/samples/sample1,sample2/info?study=1000genomes |
As it is explained later in this documentation, this RESTful web service will return the information stored in OpenCGA of the user opencga.
- apiVersion (v1) : indicates OpenCGA version to retrieve information from, data models and API may change between versions.
- resource (samples) : specifies the data type of what the user wants to retrieve. This can be one of the resources listed below.
- id (sample1,sample2) : the resource id's we want to query.
- endpoint (info) : these parameters must be specified depending on the nature of your input data. For instance, info is used to fetch the information stored in the database regarding the id's passed.
- options (study=1000genomes) : variables in key value pair form, passed as query parameters.
URL parameters
apiVersion
apiVersions are numbered as v1, v2, etc. At this moment we are heading to second stable apiVersion which will be v2.
resource
There are several metadata resources implemented such as users, samples, individuals, ... see below for more info.
IDs
This is the unique identifier(s) corresponding to the resource we want to interact with. Plural means a comma separated list of IDs can be passed to improve performance with a single REST call rather than multiple calls. OpenCGA preserves the order of the results with corresponding IDs. A Boolean variable, silent, can be set to indicate, in case of a failure (resource doesn't exist, permission denied etc), whether the user is interested in receiving partial results (true) with the information that could be successfully retrieved or just a failure with no results. As a trade off between performance and ease of use a maximum of 100 IDs are allowed in one web service.
options
These query parameters can modify the behaviour of the query (exclude, include, limit, skip and count) or add some filters to some specific endpoints to add useful functionality. The following image shows some typical options for a certain web service.
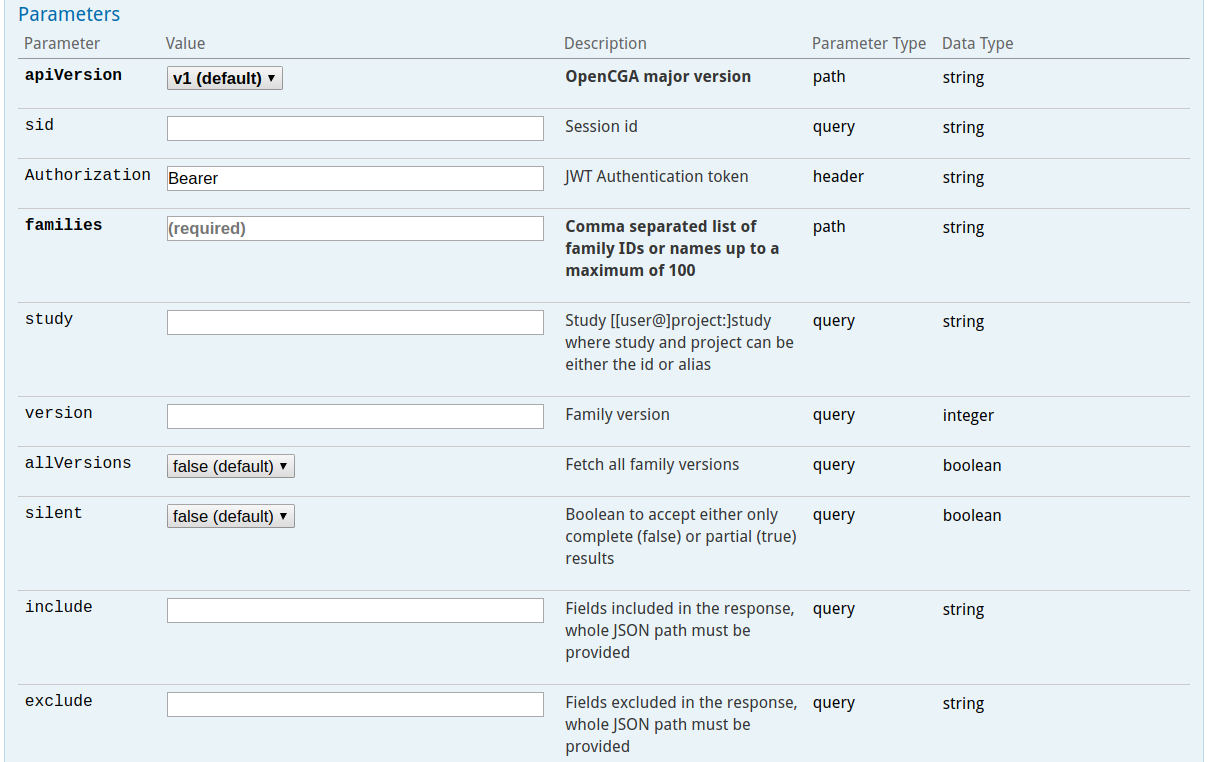
REST Response
OpenCGA 1.x
Most web services return the results encapsulated in a single QueryResponse object (view data model) consisting of some metadata and a list of QueryResult objects (view data model) called response containing the data and metadata requested. The reason for this two-level response is that some REST web services allow to pass multiple IDs as input parameter, this improves significantly the performance by reducing the number of calls, for instance a calling /info method with three sample IDs will return a QueryResponse object with three QueryResults. Then, each QueryResult can contain multiple results, for instance when getting all samples from an individual or when fetching all variants from a gene.
The Response object as well as the behaviour regarding the nested lists change in OpenCGA 2.x (see below) |
However, most of the web services will return a QueryResponse with one single QueryResult with one or more result. In general the response object looks like:
{
"apiVersion": "v1",
"time": 19,
"warning": "",
"error": "",
"queryOptions": {
"metadata": true,
"skipCount": false,
"limit": 10
},
"response": [
{
"id": "search",
"dbTime": 18,
"numResults": 10,
"numTotalResults": 56,
"warningMsg": "",
"errorMsg": "",
"resultType": "",
"result": [
{
// result 1
},
{
// result 2
},
// ...
{
// result 10
}
]
}
]
} |
where:
- Line 1: single QueryResponse object
- Lines 2 and 3: show the version and the duration time (ms)
- Lines 4 and 5: show warning and error messages, for instance when having network issues you could get "Catalog database not accessible"
- Line 6: summary of all option parameters provided
- Line 11: list of QueryResults called response. In this example, and in most of calls, there is only one QueryResult.
- Line 14: database duration time (ms) for each QueryResult.
- Line 15 and 16: number of elements returned in the list result (see below) and total number of records found in the database for a given query.
- Line 17 and 18: specific warning and error messages for each QueryResult
- Line 19: type of result such as resource.
- Line 20: list of results for this query, this can be samples, variants, ...
OpenCGA 2.x
Most web services return the results encapsulated in a single RestResponse object (view data model) consisting of some metadata and a list of DataResult objects (view data model) called responses containing the data and metadata requested. The first response of the list will always contain the response of the OpenCGA federation being directly queried. Any additional response in the list will belong to other federated servers that could be connected. Each federated response will contain a list of results (DataResults) containing the data that has been queried.
{
"apiVersion": "v2",
"time": 23,
"params": {
"include": "id",
"study": "study1",
"limit": "3"
},
"events": [
{
"type": "WARNING",
"message": "This is a development version OpenCGA 2.0.0-RC"
}
],
"responses": [
{
"time": 16,
"events": [],
"numResults": 3,
"results": [
{
"id": "HG01879"
},
{
"id": "HG01880"
},
{
"id": "HG01881"
}
],
"resultType": "org.opencb.opencga.core.models.Sample",
"numMatches": 3502,
"numInserted": 0,
"numUpdated": 0,
"numDeleted": 0
}
]
} |
where:
- Line 1: single RestResponse object
- Lines 2 and 3: show the version and the duration time (ms)
- Lines 4-8: show all the parameters that have been provided.
- Line 9-14: show an events array where info, warning and error messages will be shown: For instance, when having network issues you could get "Catalog database not accessible".
- Line 15: list of DataResults called responses. In this example, because federation is disabled, it only contains a single DataResult.
- Line 17: database duration time (ms) for each DataResult.
- Line 18: list of events where info, warning and error messages will be shown. For instance, it can show messages such as "Permission denied to access sample xxx".
- Line 19: number of elements returned in the results list.
- Line 20-30: List of results for this query.
- Line 31: resource type of results.
- Line 32: total number of records found in the database for the given query.
- Line 33-35: Number of elements inserted, updated and deleted in the database. These counters only make sense for create, updated and delete operations.
Resources and Endpoints
REST API is organised into two main groups of web services, one to work with metadata and a different one to run some analyses: Catalog and Analysis. See below a description of the web services.
Catalog Web Services
Contains all endpoints for managing and querying metadata and permission.
| Resource | Path | Description | Main Endpoints |
|---|---|---|---|
| Users | /users | Different methods to work with users | info, create, login, ... |
| Projects | /projects | Projects are defined for each user and contains studies | info, create, studies, ... |
| Studies | /studies | Studies are the main component of OpenCGA Catalog. They can be shared with other users and are the containers of the data (files, samples, cohorts, jobs...). | info, create, groups, ... |
| Files | /files | Files are added to the study and can be indexed to be queried | info, create, index, share, ... |
| Jobs | /jobs | Jobs are used to execute analyses. | info, create, ... |
| Families | /families | Family is a connected collection of individuals based on their relationship. | info, create, ... |
| Individuals | /individuals | Individual is the member from which a sample was taken. | info, create, ... |
| Samples | /samples | Samples are each of the experiment samples, typically matches a NGS BAM file or VCF sample. | info, create, annotate, share, ... |
| Cohorts | /cohorts | Cohort is a group of samples that share some common properties. These are used for data analysis. | info, create, stats, samples, ... |
| Clinical Analysis | /clinical | This handles creating and search of a clinical analyses. | info, create, ... |
| Meta | /meta | Contains basic information about the status of an OpenCGA installation instance. | ping, about, status |
| GA4GH | /ga4gh | GA4GH standard web services to search genomics data in OpenCGA | variant search, reads search, responses |
Analysis Web Services
Different endpoint for running the alignment, variant and clinical analysis
| Category | Path | Description | Main Endpoints |
|---|---|---|---|
| Alignment Analysis | /analysis/alignment | Operations over Read Alignments to facilitate complete analysis with different tools. | index, query, stats, coverage |
| Variant Analysis | /analysis/variant | Operations over Genomic Variants to facilitate complete analysis with different tools. | index, stats, query, validate, ibs, facet, samples, metadata |
| Clinical Analysis | /analysis/clinical | You can manage Clinical Analysis metadata (e.g create a case, set permissions) or run a genome interpretation | execute |
Swagger
OpenCGA has been documented using Swagger project. Detailed information about resources, endpoints and options is available at:
http://bioinfo.hpc.cam.ac.uk/opencga-demo
Client Libraries
Currently OpenCGA implements the following four client libraries:
Deprecation Policy
Certain APIs are deprecated over the period of time as OpenCGA is a live project and continuously improved and new features are implemented. Deprecation cycle consists of a warning period to let make user aware that these services are considered for change and highly likely will be replaced followed by a deprecated message. OpenCGA supports deprecated services for two releases (Deprecated and Next one). Deprecated services are hidden from Swagger in the following release and completely removed in the next one.
Warning (working) ---> Deprecated (working) ---> Hidden (working) ---> Removed (not working) |
All deprecated web services are clearly marked as deprecated along with a warning help message for user.
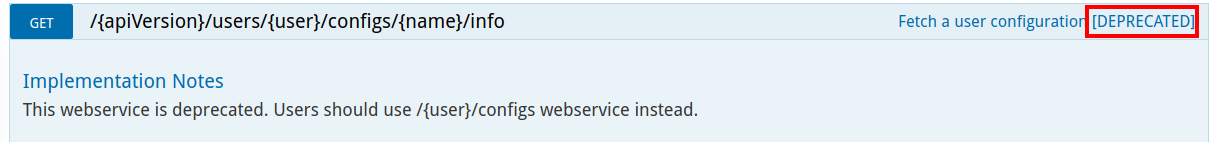
Table of Contents: