Genomic and Clinical Data
For this proof of concept (PoC) we loaded all the genomic variants of about 4,700 genomes from Genomics England, variants were loaded and indexed in the development version OpenCGA 2.0.0-beta. In total we loaded about 200 million unique variants from 4,700 gVCF files accounting for about 20TB of compressed disk space. It is worth noting that these files were generated using Dragen 2.x and the are unusually big, about 5-6GB per file.
Platform
For this proof of concept (PoC) we used the development version OpenCGA v2.0.0-beta using the Hadoop Variant Storage Engine that uses Apache HBase as back-end. We also used CellBase 4.6 for the variant annotation.
For the platform we used a 10-nodes Azure HDInsight 3.6 cluster using Data Lake Storage Gen2. HDInsight 3.6 uses Hortonworks HDP 2.6.5 (with Hadoop 2.7.3 and HBase 1.1.2) and we used Azure Batch for loading concurrently all the VCF files which had been copied previously to a NFS server, you can see details here:
| Node Type | Nodes | Azure Type | Cores | Memory (GB) | Storage |
|---|---|---|---|---|---|
| Hadoop Master | 3 | Standard_D12_V2 | 4 | 28 | Data Lake Gen2 |
| Hadoop Worker | 10 | Standard_DS13_V2 | 8 | 56 | Data Lake Gen2 |
| Azure Batch Queue | 20 | Standard_D4s_v3 | 4 | 16 | NFS Server |
Initially, we evaluated the new HDInsight 4.0, although it worked quite well there were some few minor issue, so for this PoC we decided to use the more stable HDInsight 3.6 (HDI3.6) over Data Lake Gen2 (DL2), we will refer to this as HDI3.6+DL2. During the PoC we worked with Azure engineers to debug and fix all these issues, unfortunately we did no have time to repeat the benchmark.
As you will see below in the Analysis Benchmark section, once we completed the PoC we increased the size of HDInsight to 20 nodes and repeated some tests to study the performance improvement.
Genomic Data Loading and Indexing
In order to study the loading performance we set up a Azure Batch queue of 20 computing nodes. This allowed us to load multiple files at the same time from different servers. We configured Azure Batch to load 1 VCF file per node resulting in 20 files being loaded in HBase simultaneously. During the load we studied two different configurations:
- Loading VCF files vs. loading transformed files
- Using HDD vs. SSD disk in the NFS server
You can observe the results in the following plot:
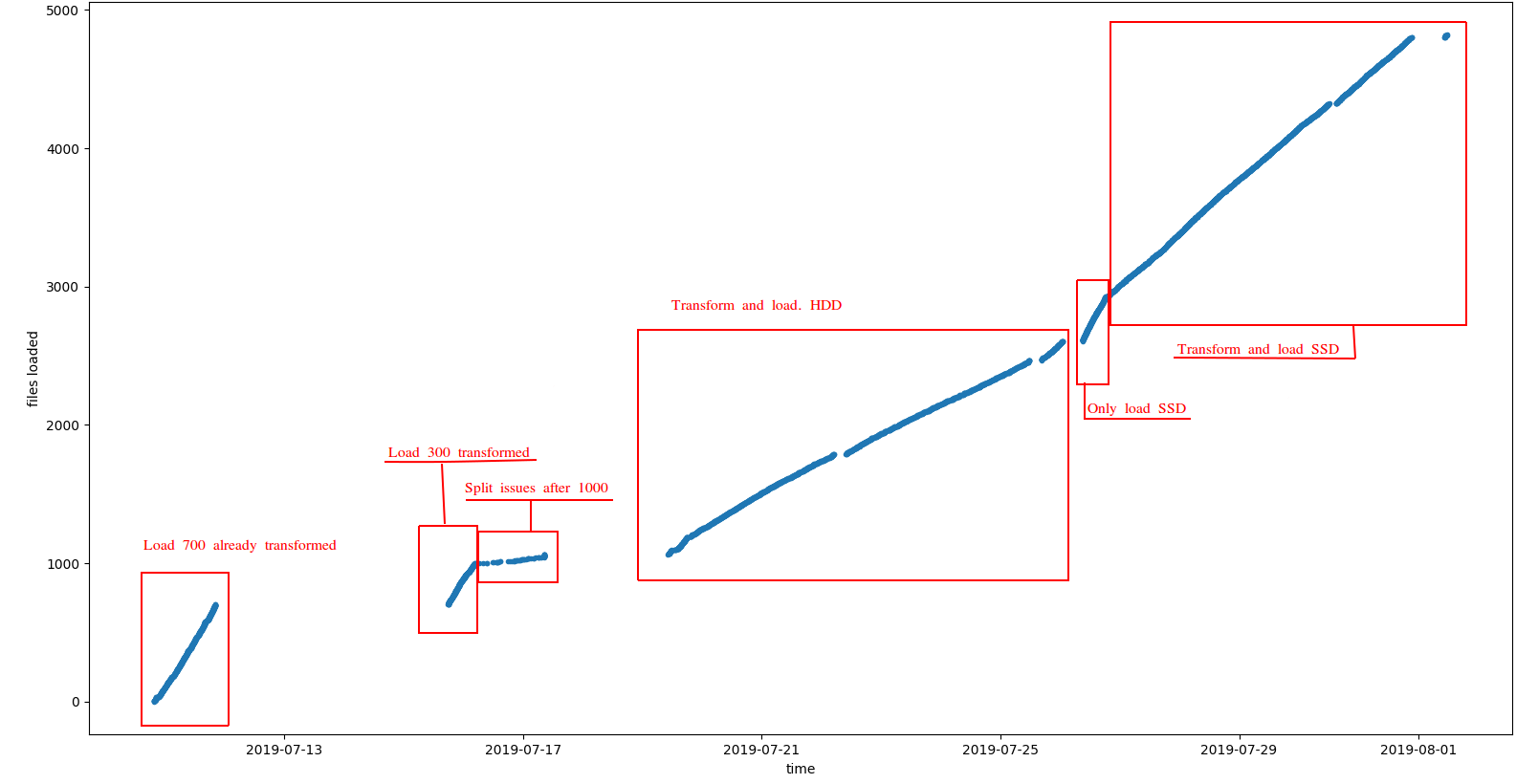
Some comments:
- As expected loading already transformed files is much faster since we only need to load and index data in HBase. See this link for more information Indexing Genomic Variants
- Also, loading from SSD disks showed a better performance
The most typical scenario when indexing genomic data is to transform+load at the same time, so assuming SSD disk the observed performance was about 380 VCF files indexed a day, or about 2TB/day. It is worth noting that:
- gVCF used were several times bigger than usual
- the number of Hadoop worker nodes was just 10
- we loaded up to 20 files concurrently but this could have been increased
These variables have a huge impact in the indexing performance, the expected performance with more real gVCF files and production cluster is more than 1,000 VCF a day.
The more representative section is the last one, where we upgraded the input disk to speed up the reading. In average, with the improved disk, processing up to 20 files simultaneously we have these numbers:
| Time | Time/nodes | |
|---|---|---|
| Transform | 00:29:36 | 00:01:28 |
| Load | 00:46:19 | 00:02:19 |
| Total | 1:15:55 | 00:03:48 |
Index speed:
- 15.8 files/h
- 379.4 files/day
- 79.0 GB/h
- 1.85 TB/day
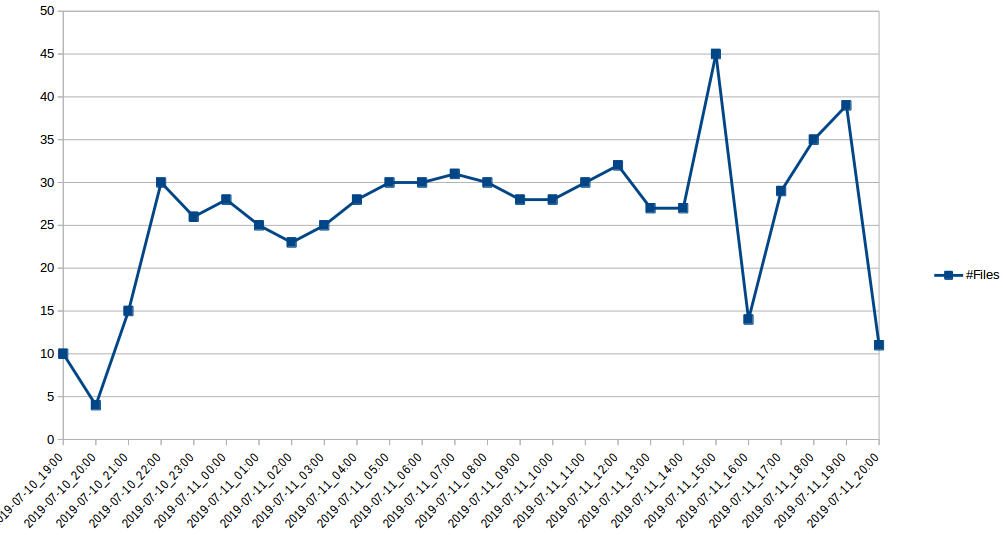
#Files Day Hour 10 2019-07-10 19 4 2019-07-10 20 15 2019-07-10 21 30 2019-07-10 22 26 2019-07-10 23 28 2019-07-11 00 25 2019-07-11 01 23 2019-07-11 02 25 2019-07-11 03 28 2019-07-11 04 30 2019-07-11 05 30 2019-07-11 06 31 2019-07-11 07 30 2019-07-11 08 28 2019-07-11 09 28 2019-07-11 10 30 2019-07-11 11 32 2019-07-11 12 27 2019-07-11 13 27 2019-07-11 14 45 2019-07-11 15 14 2019-07-11 16 29 2019-07-11 17 35 2019-07-11 18 39 2019-07-11 19 11 2019-07-11 20 |
Operations
First batch of 700 files
74.096.015 variants
Aggregate
Prepare: 529.303s [ 00:08:49 ]
Aggregate: 9591.626s [ 02:39:52 ]
Write: 7012.733s [ 01:56:53 ] -> Size : 59.5 GiB
Stats
1352.675s [ 00:22:33 ]
Annotate
Prepare: 722.327s [ 00:12:02 ]
Annot: 50384.383s [ 13:59:44 ]
Load: 28204.666s [ 07:44:04 ]
SampleIndex: 12403.542s [ 03:26:44 ]
Secondary index (Solr)
.....
Analysis Benchmark
Query and Aggregation Stats
Stats
| Operation | 10 Nodes | 20 Nodes | Speed-up |
|----------------------|-------------------------|-------------------------|----------|
| Stats | 6624.336s [ 01:50:24 ] | 3199.123s [ 00:53:19 ] | 2.07 |
| SampleIndex | 9930.483s [ 02:45:30 ] | 6400.994s [ 01:46:41 ] | 1.55 |
| SampleIndex annotate | 17593.733s [ 04:53:14 ] | 9453.761s [ 02:37:34 ] | 1.86 |
GWAS
Clinical Analysis
Table size
| Table | Compression | Size (TB) |
|---|---|---|
| Variants table | GZ | 2.9 |
| Variants table | SNAPPY | 4.7 |
Table of Contents: