Overview
This operation provides an advanced genome VCF merging to provide a mult-isample dataset suitable for cohort analysis. Along with Annotation and Statistics, this is an optional enrichment operation over the database. [#713] [#757] [#877]
This operation is designed to find a proper value for the unknown genotype values, reading the reference blocks from the gVCF files.
Given a set of samples, the process iterate over all variants where some, but not all samples have missing values (where the value is not present, not the same as the genotype ./.). A sample can have missing value in three situations:
- The sample had a Reference Block in that position in the original VCF
Actions here will be to copy the sample and file information from the block. FileEntry.call will point to the used reference block. - The sample had a different variant overlapping with that position.
In this case, the alternates from the overlapping variant will be written as secondary alternate in the file column, and the INFO and FORMAT information, reordered. FileEntry.call will point to the overlapping variant. - The sample has no record for that position in the original VCF.
This situation can only happen if there is an error in the gVCF, or if the fill-gaps was done for VCF files. The genotype will be filled with?/?and the rest of values with missing.
Executing this operation against all the samples in the database can be really expensive in terms of time and disk usage, because it will fill all the gaps in the spare matrix that the variants table is. To avoid this situation, this operation skips the samples where the genotype is homozygous reference (HOM_REF, 0/0), and the files where all the belonging samples are HOM_REF.
This Operation is only available in HBase Storage Engine in Hadoop |
Family-based Aggregation
This operation is slightly different from the general aggregation. It is designed to work only with a family, and will write all the genotypes, even the HOM_REF, and the related sample data that validates the genotype.
Example
In the next figure we can see an example of aggregating multiple variants, from different single-sample files.
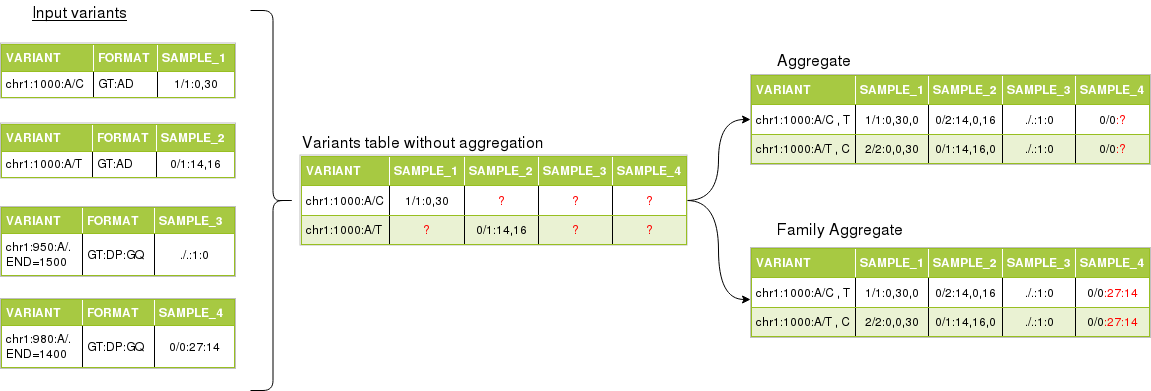
The variants from samples 1 and 2 have two overlapping variants. Variants from samples 3 and 4 are reference blocks from a gVCF.
Special scenarios
There are some scenarios where the result of the aggregation operation is not obvious, and should be defined and handled carefully:
Multiple overlaps
This scenario consists of having multiple overlapping positions in one variant. This may happen because of many reasons:
- Deletion from sample A overlapping with N smaller variants from sample B
- Inconsistent input VCF with overlapping variants
In any of this two situations, we can not determine genotype of the sample B in the deletion. We should mark that there is something in this position, so we can not mark as ./. or 0/0. For this, we should use the special allele<*>from the VCF spec v4.3 (known as<NON_REF>at GATK). The sample B will have the genotype<*>/<*>(i.e.2/2where<*>is the second alternate) for the deletion.
- Deletion from sample A overlapping with N reference blocks from sample B
Overlap with a split multi-allelic variant
In this scenario, a variant from file A may overlap with many variants produced from the split of a multi-allelic variant from file B. The information in these split variants from B is the same (just rearranged), so we know exactly what is in this position. All the overlapping variants share the same
FileEntry.call, which contains the original call of this variant. We should just take any of them.
Structural variants
Incomplete gVCF file
Insertion not overlapping with any variant.
Table of Contents: