Overview
Indexing variants does not apply any modification to the generic pipeline. The input file format is VCF, accepting different variations like gVCF or aggregated VCFs
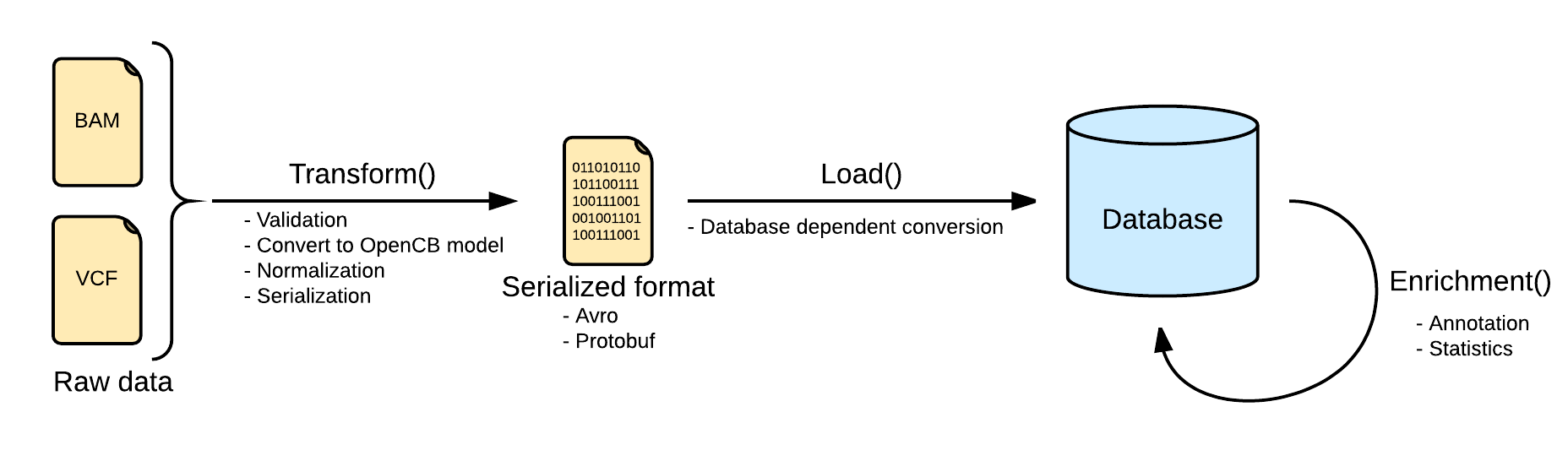
Transform
Files are converted Biodata models. The metadata and the data are serialized into two separated files. The metadata is stored into a file named <inputFileName>.file.json.gz serializing in json a single instance of the biodata model VariantSource, which mainly contains the header and some general stats. Along with this file, the real variants data is stored in a file named <inputFileName>.variants.avro.gz with a set of variant records described as the biodata model Variant.
VCF files are read using the library HTSJDK, which provides a syntactic validation of the data. Further actions on the validation will be taken, like duplicate or overlapping variants detection.
By default, malformed variants will be skipped and written into a third optional file named <inputFileName>.malformed.txt . If the transform step generates this file, a curation process should be taken to repair the file. Otherwise, the variants would be skipped.
All the variants in the transform step will be normalized as defined here: Variant Normalization. This will help to unify the variants representation, since the VCF specification allows multiple ways of referring to a variant and some ambiguities.
Load
Loading variants from multiple files into a single database will effectively merge them. In most of the scenarios, with a good normalization, merging variants is strait forward. But in some other scenarios, with multiple alternates or overlapping variants, the merge requires more logic. More information at Merge Mode.
Details about load are dependent on the implementation.
Limitations
- You can not load two files with the same sample in the same study. See OpenCGA#158.
There is an exception for this limitation for the scenarios where the variants were split in multiple files (by chromosome, by type, ...). In this case, you can use the parameter--load-split-data. SeeOpenCGA#696 - You can not index two files with the same name (e.g. /data/sample1/my.vcf.gz and /data/sample2/my.vcf.gz) in the same study. This limitation should not be a problem in any real scenario, where every VCF file usually has a different name. If two files have the same name, the most likely situation is that they contain the same samples, and this is already forbidden by the previous limitation.
Table of Contents: