Versions Compared
Key
- This line was added.
- This line was removed.
- Formatting was changed.
One of the goals of The 100,000 Genomes Project from Genomics England is to enable new medical research. Researchers will study how best to use genomics in healthcare and how best to interpret the data to help patients. The causes, diagnosis and treatment of disease will also be investigated. This is currently the largest national sequencing project of its kind in the world.
To achieve this goal Genomics England set up a Research environment for the researchers and installed OpenCGA, CellBase and IVA from OpenCB. We have loaded 64,078 whole genomes in OpenCGA, in total more than 1 billion unique variants have been indexed in the OpenCGA Variant Storage, and all metadata and clinical data for samples and patients have been loaded in OpenCGA Catalog. Variants were annotated using CellBase and IVA front-end was installed to analyse and visualise the data. Here you can find a full report of about the loading and analysis of the 64,078 genomes.
Genomic and Clinical Data
The variants and clinical data of 64,078 participant genomes been loaded and indexed in OpenCGA. In total they represent more than 30,000 VCF files compressed accounting for about 40TB of disk space. This data is divided in four different datasets depending on the genome assembly (GRCh37 or GRCh38) and the type of study (germline or somatic), this data has been organised in OpenCGA in three different Projects and four Studies:
| Project | Study ID and Name | Samples | VCF Files | VCF File Type | Samples/File | Variants |
|---|---|---|---|---|---|---|
| GRCh37 Germline | RD37 Rare Disease GRCh37 | 12,142 | 5,329 | Multi sample | 2.28 | 298,763,059 |
| GRCh38 Germline | RD38 Rare Disease GRCh38 | 33,180 | 16,591 | Multi sample | 2.00 | 437,740,498 |
CG38 Cancer Germline GRCh38 | 9,167 | 9,167 | Single sample | 1.00 | 286,136,051 | |
| GRCh38 Somatic | CS38 Cancer Somatic GRCh38 | 9,589 | 9,589 | Somatic | 1.00 | 398,402,166 |
| Total | 64,078 | 40,676 | 1,421,041,774 | |||
As you can see each dataset is loaded in OpenCGA as a different Study, an these are grouped in three different Projects depending on the type of data (germline or somatic) and assembly (GRCh37 or GRCh38).OpenCGA Catalog stores all the metadata and clinical data of files, samples, individuals and cohorts. Rare Disease studies also include pedigree metadata by defining families. Also, a Clinical Analysis has been defined for each family. Several Variable Sets have been defined to store GEL custom data for all these entities.
Platform
For the Research environment we have used OpenCGA v1.4 using the new Hadoop Variant Storage that use Apache HBase as back-end because of the huge amount of data and analysis needed. We have also used CellBase v4.6 for the variant annotation. Finally we set up a IVA v1.0 web-based variant analysis tool.
The Hadoop cluster consists of about 30 nodes running Hortonworks HDP 2.6.5 (with HBase 1.1.2) and a LSF queue for loading all the VCF files, see this table for more detail:
| Node | Nodes | Cores | Memory (GB) | Storage (TB) |
|---|---|---|---|---|
| Hadoop Master | 5 | 28 | 216 | 7.2 (6x1.2) |
| Hadoop Worker | 30 | 28 | 216 | 7.2 (6x1.2) |
| LSF Loading Queue | 10 | 12 | 364 | Isilon storage |
OpenCGA 1.4.2 with the Hadoop Variant Storage has been used.
Loading Data
The data ingestion is executed in the LSF nodes, connected directly to the Hadoop platformGenomic Data Load
In order to improve the load performance we set up a small LSF queue of ten computing nodes. This configuration allows us to run load multiple files at the same time. Having 10 worker nodes in the queue, each of them loading We configured LSF to load up to 6 files at the same time, results VCF files per node resulting in 60 files being loaded concurrently.
Multi sample filesin HBase in parallel without any incidence, by doing this we observed a 50x in loading throughput. This resulted in an average of 125 VCF files loaded per hour in studies RD37 and RD38, about 2 files per minute. In the study CG38 the performance was 240 VCF files per hour or about 4 files per minute.
RD37 and RD38
The files from Rare Disease studies (RD38 & RD37) contain more than one sample per file. In average, 2 samples per file. This results in larger files, increasing the loading time, compared with single-sample files.
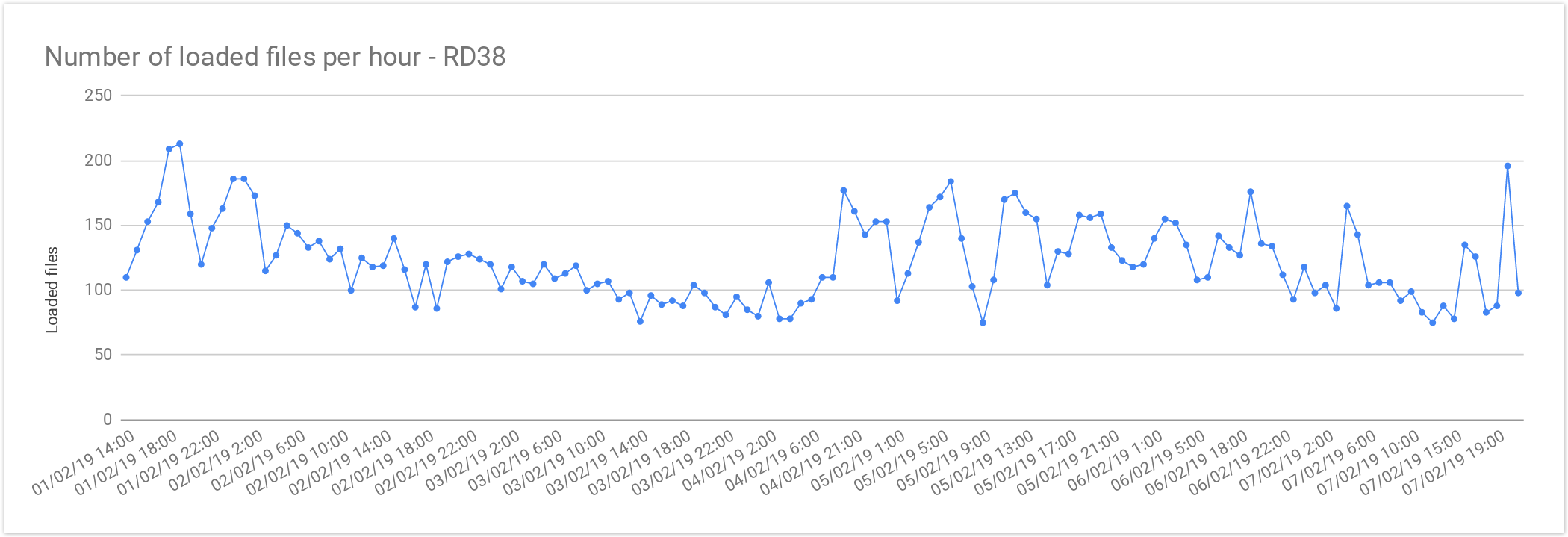
| Concurrent files loaded | 60 |
|---|---|
| Average files loaded per hour | 125.72 |
| Load time per file | 00:28:38 |
Saturation Study
As part of the data loading process we decided to study the number of unique variants added in each batch of 1,000 samples. We generated this saturation plot for RD38:
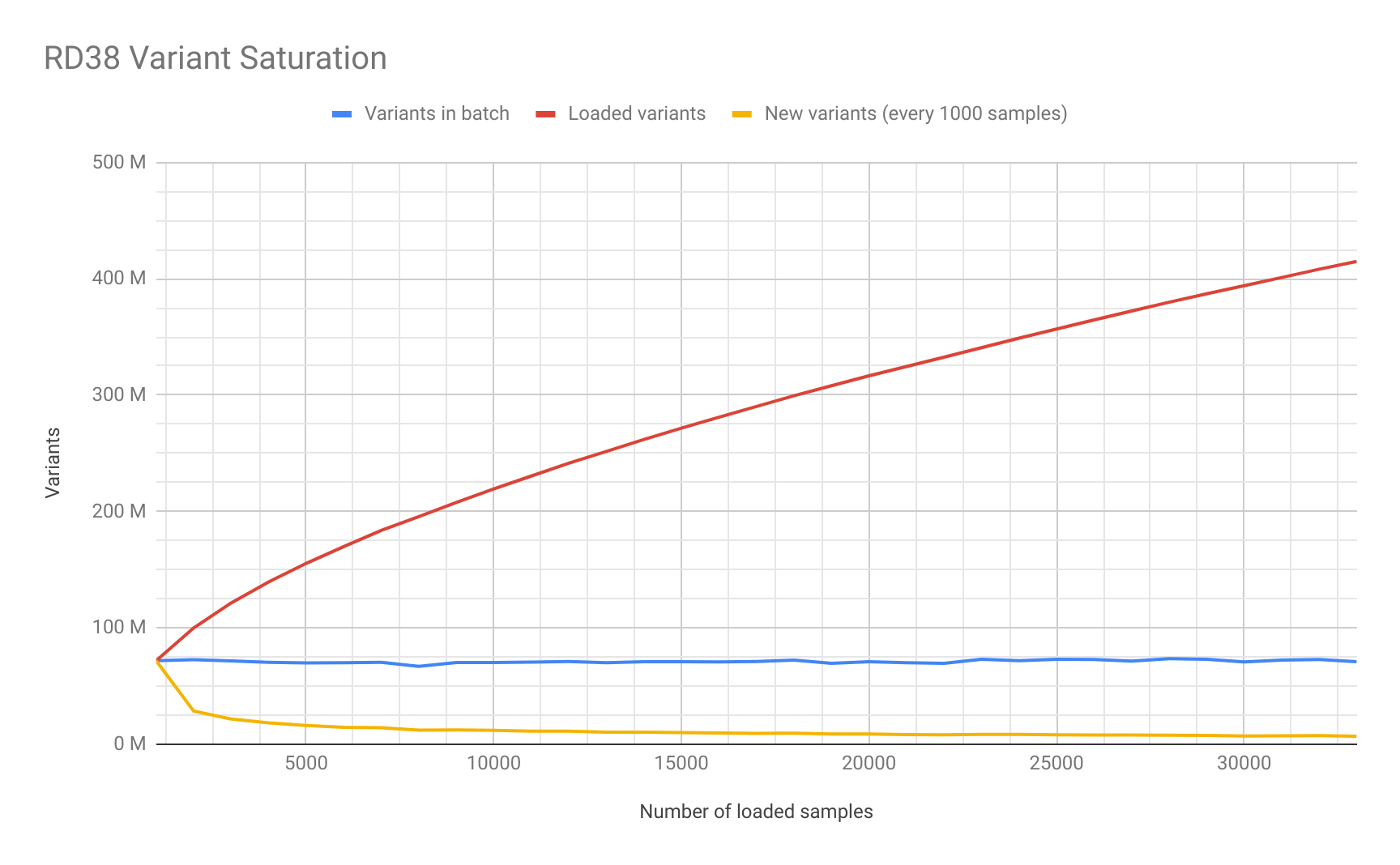 Image Added
Image Added
CG38
The files from Cancer Germline studies (CG38) contain one sample per file. Compared with the Rare Disease, these files are smaller in size, therefore, the load is slightly faster.
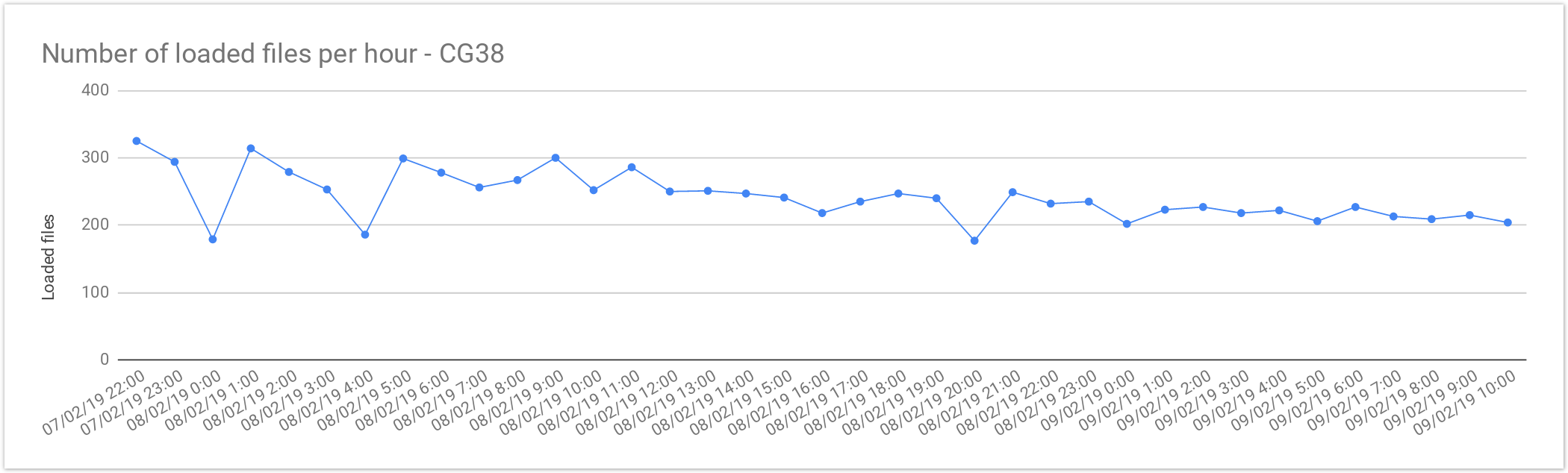
| Concurrent files loaded | 60 |
|---|---|
| Average files loaded per hour | 242.05 |
| Load time per file | 00:14:52 |
Analysis Benchmark
We would like to distinguish two types of queries: General and Clinical
General QueriesQuery and Aggregation Stats
These queries are are only filtering by variant annotation and cohort stats. These queries only include aggregated data, not returning sample genotypes.
| Filter | Results | Total Results | Time |
|---|---|---|---|
| consequence type = LoF + missense_variant | 10 | 3704626 | 0.189s |
consequence type = LoF + missense_variant biotype = protein_coding | 10 | 3576472 | 0.260s |
panel with 200 genes | 10 | 3882902 | 0.299s |
Clinical
QueriesAnalysis
Clinical queries, or sample queries, enforces queries to return variants of a specific set of samples. These queries can use all the filters from the general queries. The result will include a ReportedEvent for each variant, which determines possible conditions associated to the variant.
| Filter | Results | Total Results | Time |
|---|---|---|---|
Segregation mode = biallelic filter = PASS | 10 | 211787 | 0.420s |
Segregation mode = biallelic filter = PASS | 2000 | 211787 | 1.079s |
De novo variants filter = PASS consequence type = LoF + missense_variant | 24 | 24 | 0.680s |
| Compound Heterozygous filter = PASS biotype = protein_coding consequence type = LoF + missense_variant | 717 | 717 | 10.995s |
User Interfaces
Python and R
Acknowledgements
We would like to thank Genomics England very much for their support and for trusting OpenCGA and the rest of OpenCB suite for this amazing release.
Table of Contents:
| Table of Contents | ||
|---|---|---|
|