Versions Compared
compared with
Key
- This line was added.
- This line was removed.
- Formatting was changed.
Here you can find a full report of about loading 62,000 samples for Genomics England Research environment.
Platform
The platform used for this case study consists on a Hadoop Cluster of 35 nodes (5 + 30) and a LSF queue system:
| Node | #nodes | cores | memory (GB) |
|---|---|---|---|
| LSF queue node for load | 10 | 12 | 364 |
| Hadoop master nodes | 5 | 28 | 216 |
| Hadoop worker nodes | 30 | 28 | 216 |
Data
The data of this case study contains a total of 64,078 samples divided in 4 different datasets.
| Dataset | Alias | Files | File type | Samples | Samples per file | Variants |
|---|---|---|---|---|---|---|
| Rare Disease GRCh38 | RD38 | 16,591 | Multi sample VCF | 33,180 | 2.00 | 437,740,498 |
| Cancer Germline GRCh38 | CG38 | 9,167 | Single sample VCF | 9,167 | 1.00 | 286,136,051 |
| Cancer Somatic GRCh38 | CS38 | 9,589 | Somatic VCF | 9,589 | 1.00 | 398,402,166 |
| Rare Disease GRCh37 | RD37 | 5,329 | Multi sample VCF | 12,142 | 2.28 | 298,763,059 |
| Total | 40,676 | 64,078 | 1,421,041,774 | |||
Each dataset is loaded in OpenCGA as a study.
Loading Data
Multi sample files
The files from Rare Disease studies (RD38 & RD37) contain more than one sample per file. In average, 2 samples per file.
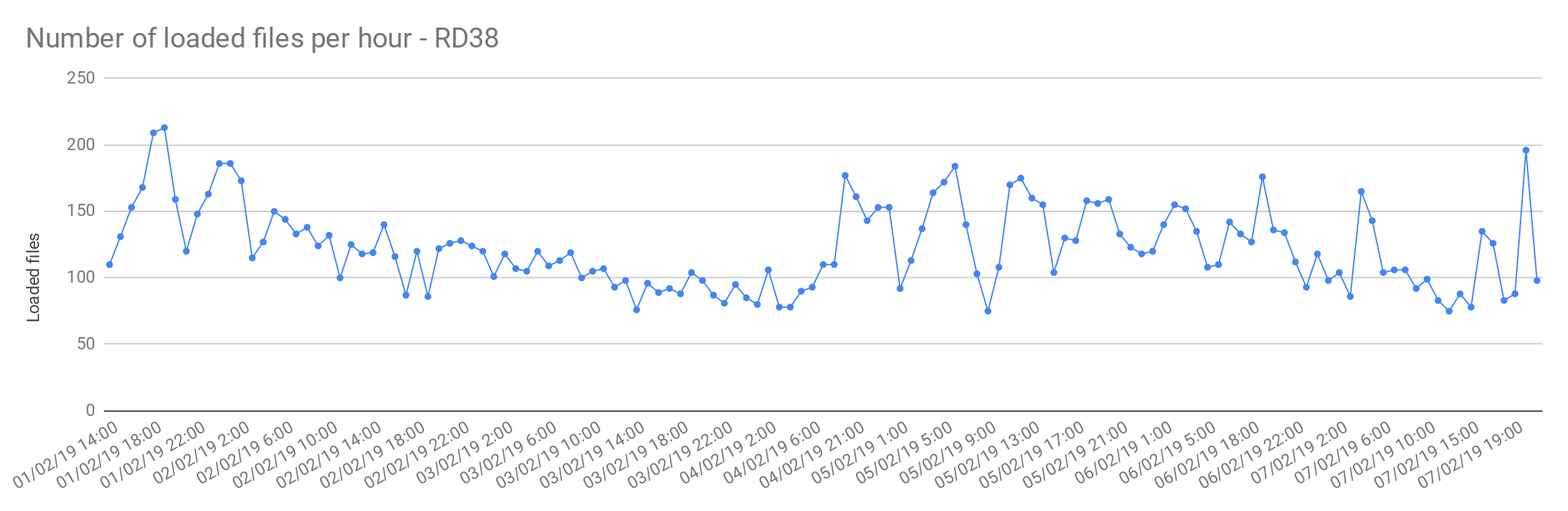 Image Added
Image Added
Single sample files
The files from Cancer Germline studies (CG38) contain one sample per file. Compared with the Rare Disease, these files are smaller in size, therefore, the load is slightly faster.
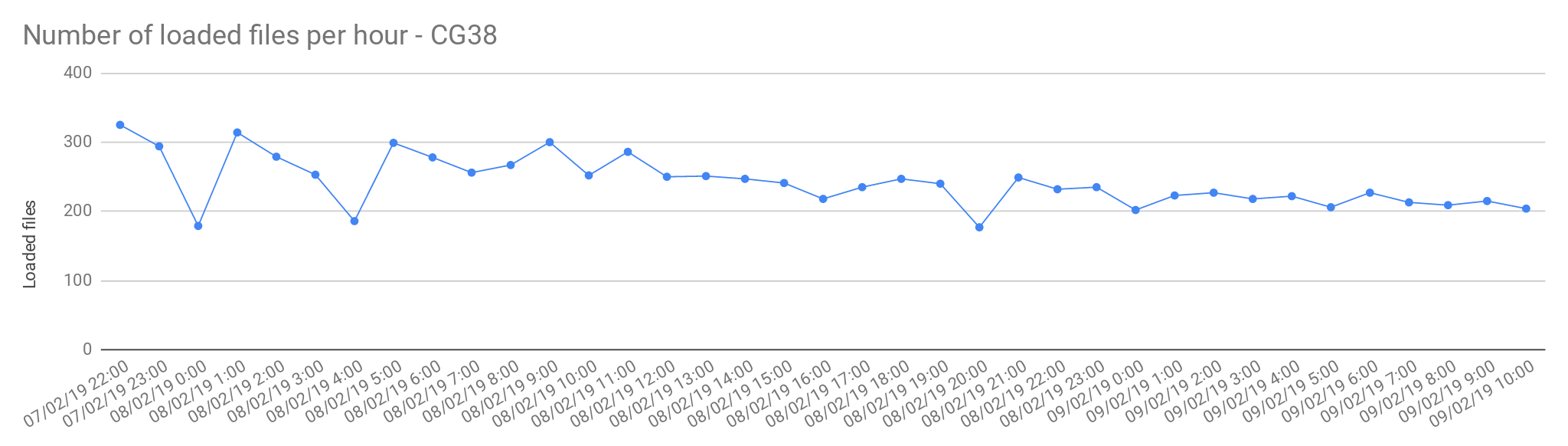 Image Added
Image Added
Query Performance
Common Queries
Clinical Queries
Table of Contents:
| Table of Contents | ||
|---|---|---|
|