Versions Compared
Key
- This line was added.
- This line was removed.
- Formatting was changed.
OpenCGA benchmark is a rich benchmark suite for storage engines supported with OpenCGA, namely mogodb and hbase. Please find below the list and detailed explanation of different components of OpenCGA Benchmark and how they work together to create benchmark.
Execution Mode
Benchmark supports the following execution mode :
- Fixed
- Random
Fixed Mode
Its a fixed set of queries written in a YML file, benchmark will take each query (default) or a selection of queries passed as IDs arguments in --query, -q option and execute these as a certain number of users (-c, - -concurrency) for a specific number of time (-r, --num-repetition). Parameters listed under baseQuery section will be applied to each individual query and can be overwritten from main query or using command line option (-B, - -baseQuery) . A sample of fixed query file is displayed below:
| Code Block | ||
|---|---|---|
| ||
---
baseQuery :
summary : true
queries :
- id : "RegionAndBiotype"
description : "Purpose of this query"
query :
region : "22:16052853-16054112"
gene : "BRCA2"
biotype : "coding"
populationFrequencyMaf : "1kG_phase3:ALL>0.1"
tolerationThreshold : 300
- id : "Region"
description : "Purpose of this query"
query :
region : "22:16052853-16054112"
tolerationThreshold : 400
.....
sessionIds :
- ""
- "" |
Following command will execute ALL queries written in fixedQueries.yml file as 10 users, five times each on REST server specified in "storage-configuration.yml" :
| Code Block | ||||
|---|---|---|---|---|
| ||||
opencga-storage-admin.sh benchmark variant --concurrency 10 --num-repetition 5 --mode FIXED --connector REST |
Complete list of options, default values and explanations can be displayed using - - help option from benchmark script :
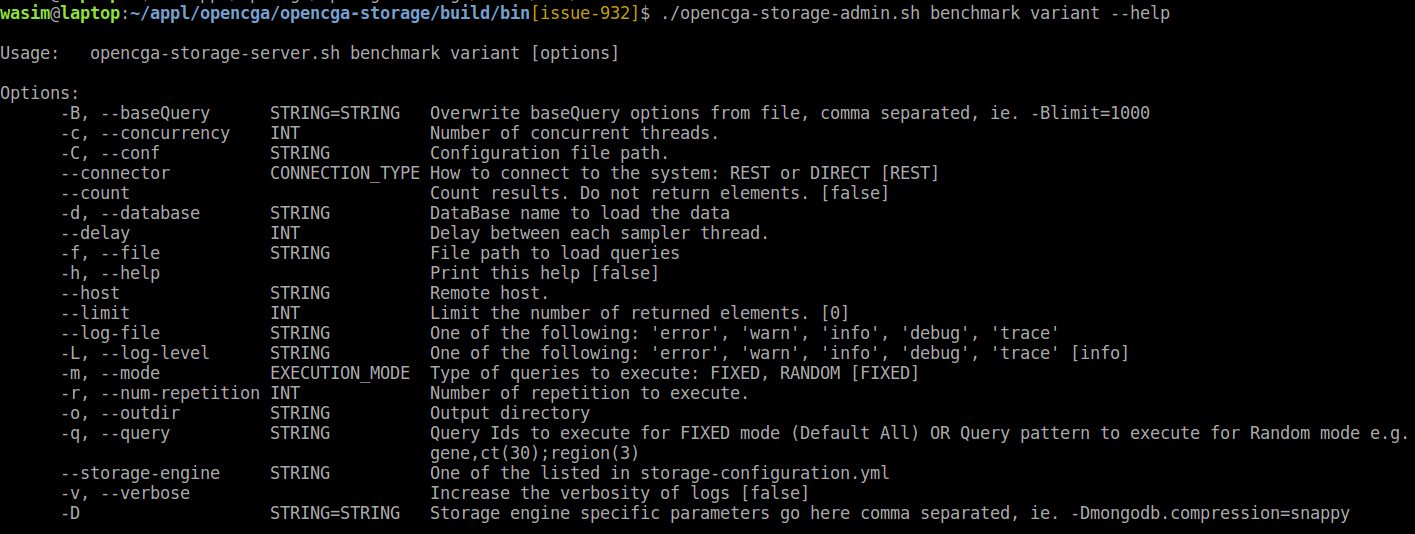
Random Mode
Random mode supports creation of random queries from meta data provided in "randomQueries.yml" and execute these on selected storage engine :
| Code Block | ||
|---|---|---|
| ||
---
baseQuery :
summary : true
exclude : studies
regions :
- chromosome : "1"
start : 1
end : 249250621
gene :
- DKFZP434A062
- GPSM1
ct : []
type :
- "SV"
- "CNV"
study :
- "1kG_phase3"
...
functionalScore :
- id : "cadd_raw"
min : 0
max : 1
- id : "cadd_scaled"
min : -10
max : 40
populationFrequencies :
- id : "1kG_phase3:ALL"
min : 0
max : 0.2
- id : "1kG_phase3:AFR"
min : 0
max : 0.15
proteinSubstitution :
- id : "polyphen"
min : 0.1
max : 0.9
operators : [">", "<"]
- id : "sift"
min : 0.1
max : 0.9
qual :
id : "polyphen"
min : 1
max : 9
operators : [">"]
conservation :
id : "phylop"
min : 0
max : 1
operators : ["=", "!="]
sessionIds :
- ""
- "" |
Following command will generate two queries one with two different "ct" values and a gene value and second with a region value provided in "randomQueries.yml" file and execute as 10 users, five times each on REST server:
| Code Block | ||
|---|---|---|
| ||
opencga-storage-admin.sh benchmark variant --concurrency 10 --num-repetition 5 --mode RANDOM -q "ct(2),gene;region" |
Storage Engine
Currently OpenCGA supports the following storage engines :
- Mongo
- HBase
- "Solr" (An optional support component for storage engines)
This value is read from "storage-configuration.yml" "defaultStorageEngineId" field or can be passed as argument on command line, --storage-engine. OpenCGA also supports "solr" to improve performance of certain queries for variant. By passing "summary=true/false" in baseQuery user can compare working and performance of OpenCGA with and without solr component.
Connection Type
Connection Type is the connection method of benchmark to the storage engine. OpenCGA currently supports three connection types :
- REST (OpenCGA web server, sessionIds are mandatory to connect using this)
- Storage REST (REST server provided with storage component and must be running state)
- Direct (Fetching data directly from storage engine using java storage adaptors)
REST type is most relevant for end users and should be used to create benchmark, remaining two are mainly to get deeper insight into the performance of storage engine without overhead of OpenCGA catalog authorisation and authentication mechanism.
Table of Contents:
| Table of Contents | ||
|---|---|---|
|