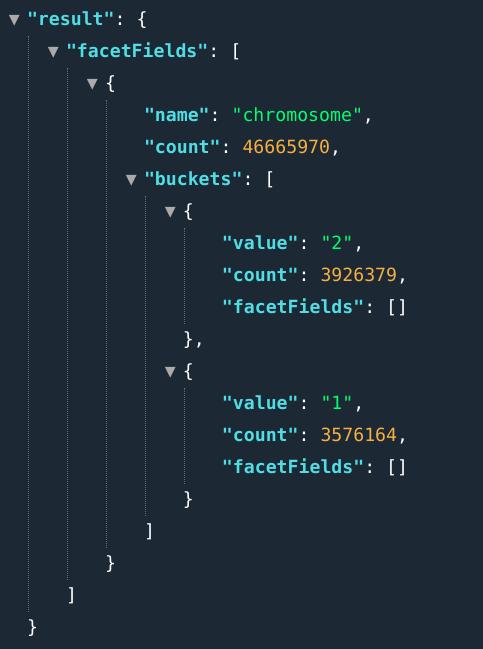
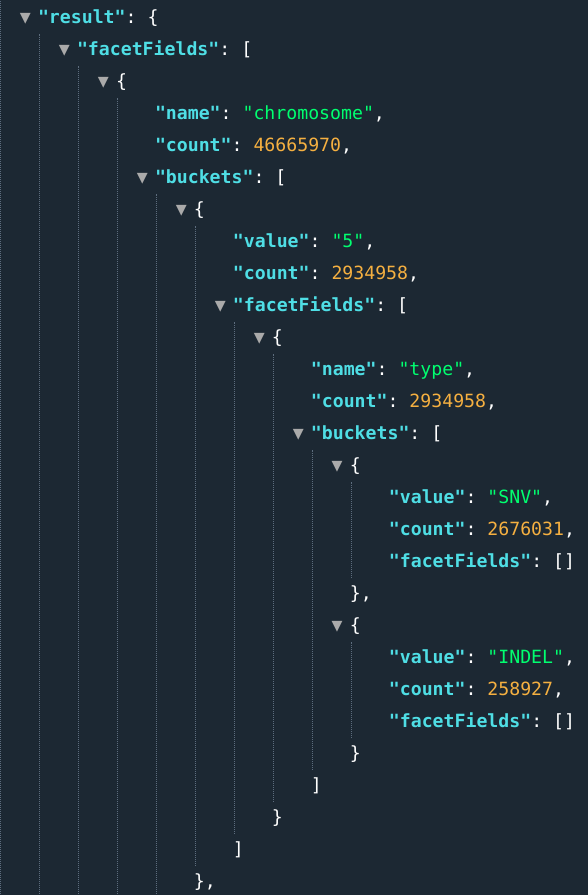
In OpenCGA, stats refers to the arrangement of search results into categories based on indexed field. Results are presented as a list of buckets, where each bucket is composed of 1) the field value and 2) a numerical count of how many matching documents were found for that field. In literature, this stats concept is known as facet or faceting as well. In fact, OpenCGA stats are based on Solr faceted search.
In addition, statsaddtion, stats allows users to query:
- Ranges that allow users to count how many documents are in an interval of a numerical field.
- Aggregation functions such as average, maximum, minimum, percentiles,...
- Nested faceted search.
...
| Code Block | ||||||
|---|---|---|---|---|---|---|
| ||||||
field_name[value1,value2,value3...]:limit |
...
| Parameter | Description |
|---|---|
field_name | The field name to produce buckets from. Mandatory. |
value1,value2,value3... | They are the values of the field name you want to select counts fromcount. They have to be enclosed in square brackets. Optional. |
limit | Number of counts to show, i.e., number of buckets. Optional. |
E.g.: ...&fields=chromosome[1,2]
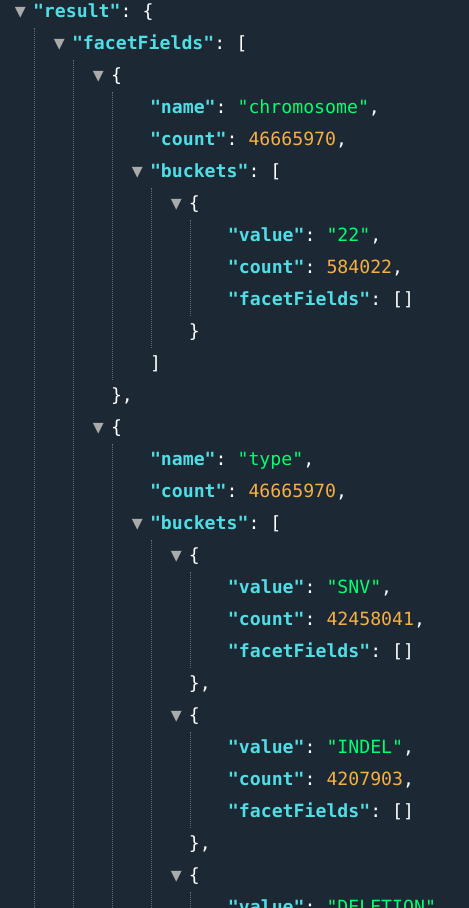
Users can query multiple stats by separating field names by semicolons.
E.g., .: ...&fields=chromosome[1,2];types
Ranges
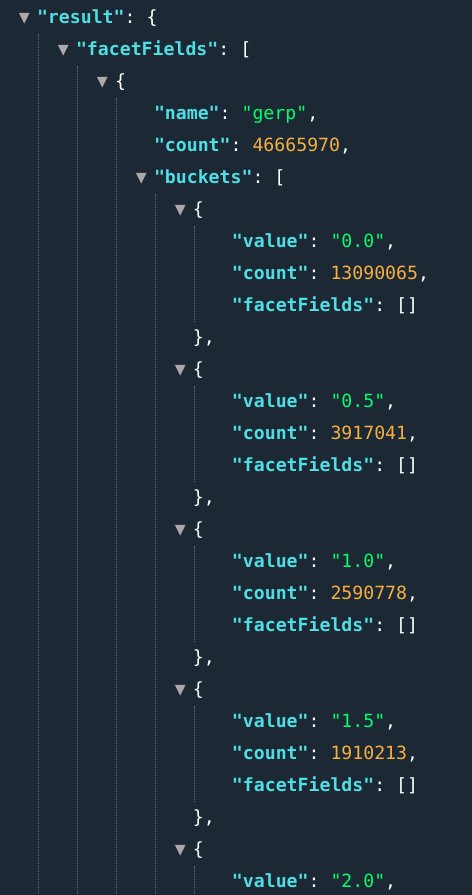
When asking for ranges, the result contains multiple buckets over a numeric field. You must specify the field name, the lower and upper bounds and the step or bucket size.
...
| Parameter | Description |
|---|---|
field_name | The numeric field name to produce range buckets from. Mandatory |
start | Lower bound of the ranges. Mandatory. |
end | Upper bound of the ranges. Mandatory. |
step | Size of each range bucket produced. Mandatory. |
E.g.: ...&fields=gerp[0..10]:0.5
Aggregation functions
Aggregation functions, also called facet functions, analytic functions, or metrics, calculate something interesting over a domain (each facet bucket).
...
| Aggregation function | Description | Example |
|---|---|---|
avg | Average of numeric values | avg(gerp) |
min | Minimum value | min(sift) |
max | Maximum value | max(caddScaled) |
unique | Number of unique values | unique(biotypes) |
hll | Distributed cardinality estimate via hyper-log-log algorithm | hll(type) |
percentile | Percentile estimates via t-digest algorithm. Calculate the percentiles: 1, 10, 25, 50, 75, 90 and 99th. | percentile(gerp) |
sumsq | Sum of squares of field or function | sumsq(caddRaw) |
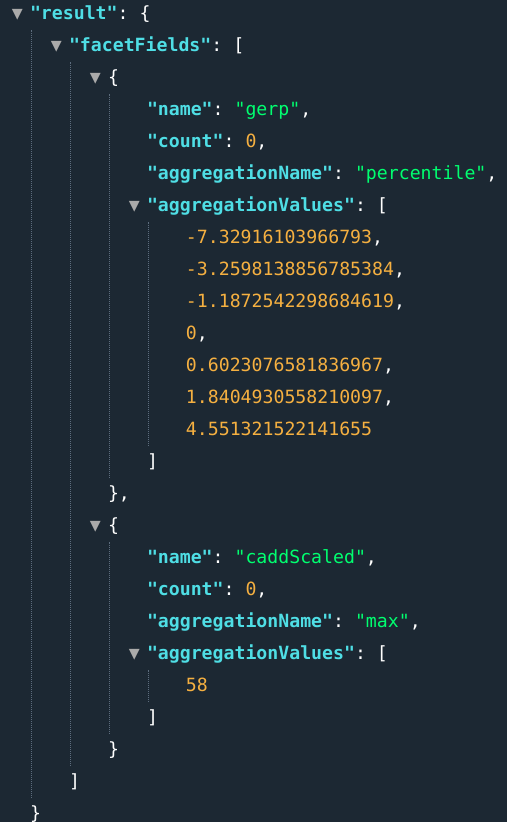
E.g., .: ...&fields=percentile(gerp);max(caddScaled)
Nested facets
Nested facets allow users to nest bucketing terms, ranges or aggregations. In order to specify nested facets you must use the symbols >>
E.g., : ...&fields=chromosome[5,6]>>type
Field types: categorical and numeric
Categorical field names
| Field name | Description | Facet example |
|---|---|---|
| id | Variant ID | List of values |
| names | Names | List of <chromosome>:<start>-<end> |
| chromosome | ||
| reference | ||
| alternate | ||
| strand | ||
| type | Variant type, e.g.: INDEL, SNV, SNP,... | List of values. Accepted values: [SNV, MNV, INDEL, SV, CNV] |
| reference | Reference | List of values |
| alternate | Allternate | List of values |
| study | Matches with variants that are in the specified studies | List of values. Accept negations. |
| file | ||
| genotype | Samples with a specific genotype e.g. HG0097:0/0;HG0098:0/1,1/1. Genotype aliases accepted: HOM_REF, HOM_ALT, HET, HET_REF, HET_ALT and MISS e.g. HG0097:HOM_REF;HG0098:HET_REF,HOM_ALT. This will automatically set 'includeSample' parameter when not provided | {samp_1}:{gt_1}(,{gt_n})*(;{samp_n}:{gt_1}(,{gt_n})*)* |
| sample | Filter variants where ALL the provided samples are mutated (HET or HOM_ALT) | List of samples. |
| filter | Specify the FILTER for any of the files. If "files" filter is provided, will match the file and the filter. | List of values. |
| qual | Specify the QUAL for any of the files. If 'file' filter is provided, will match the file and the qual | |
| info | Filter by INFO attributes from file. If no file is specified, will use all files from "file" filter. e.g. AN>200 or file_1.vcf:AN>200;file_2.vcf:AN<10 . Many INFO fields can be combined. e.g. file_1.vcf:AN>200;DB=true;file_2.vcf:AN<10 | [{file}:]{key}{op}{value}[,;]* |
| format | Filter by any FORMAT field from samples. If no sample is specified, will use all samples from "sample" or "genotype" filter. e.g. DP>200 or HG0097:DP>200,HG0098:DP<10 . Many FORMAT fields can be combined. e.g. HG0097:DP>200;GT=1/1,0/1,HG0098:DP<10 | [{sample}:]{key}{op}{value}[,;]* |
| release | Return variants that were present in that specific release | Release number |
| geneTraitId | Gene trait association ID | umls:C0007222 , OMIM:269600 |
| geneTraitName | Gene trait association name | Cardiovascular Diseases |
| clinVarTrait | ClinVar trait name | |
| gwasTrait | GWAS trait name | |
| hpo | List of HPO terms. | HP:0000545 |
| go | List of GO (Genome Ontology) terms. | GO:0002020,GO:0006508 |
| expression | List of tissues of interest | |
| proteinKeywords | List of protein variant annotation keywords | |
| drug | List of drug names |
Numeric field names
Numeric field names can be used to compute range facets and aggregation functions.
| Field name | Description | Facet range example |
|---|---|---|
| start | List of genes | |
| end | ||
| length | Consequence type SO term list. | SO:0000045,SO:0000046 |
| xref | External references | |
| biotype | List of biotypes | |
| polyphen | polyphen protein substitution score | polyphen>0.1 , sift=tolerant |
| sift | sift protein substitution score | phastCons>0.5 , phylop<0.1 , gerp>0.1 |
| phastCons | phastCons conservation score | |
| phylop | phylop conservation score | |
| gerp | gerp conservation score | |
| cadd_raw | Raw CADD functional score | |
| cadd_scaled | Scaled CADD functional score | |
| maf | Minor allele frequency | |
| populationFrequencyAlt | Alternate Population Frequency | 1000GENOMES_phase_3:AFR>0.2 |
| populationFrequencyRef | Reference Population Frequency | ESP_6500:AA<0.2 |
| populationFrequencyMaf | Population minor allele frequency | EXAC:AES>=0.6 |
| transcriptionFlags | List of transcript annotation flags | CCDS, basic, cds_end_NF, mRNA_end_NF, cds_start_NF, mRNA_start_NF, seleno |
| geneTraitId | List of gene trait association ids | umls:C0007222 , OMIM:269600 |
| geneTraitName | List of gene trait association names | Cardiovascular Diseases |
| trait | List of traits, based on ClinVar, HPO, COSMIC | |
| hpo | List of HPO terms. | HP:0000545 |
| go | List of GO (Genome Ontology) terms. | GO:0002020,GO:0006508 |
| expression | List of tissues of interest | |
| proteinKeywords | List of protein variant annotation keywords | |
| drug | List of drug names |
Variant Fields
The parameters include and exclude accepts a list of Variant Fields. This is a list with all the accepted values. Some short alias to those fields are listed in italic.
- id
- chromosome
- start
- end
- reference
- alternate
- length
- type
- studies
studies.samplesData | samples | samplesData
studies.files | files
studies.stats | stats
studies.secondaryAlternates
studies.studyId
- annotation
- annotation.ancestralAllele
- annotation.id
- annotation.xrefs
- annotation.hgvs
- annotation.displayConsequenceType
- annotation.consequenceTypes
- annotation.populationFrequencies
- annotation.minorAllele
- annotation.minorAlleleFreq
- annotation.conservation
- annotation.geneExpression
- annotation.geneTraitAssociation
- annotation.geneDrugInteraction
- annotation.variantTraitAssociation
- annotation.functionalScore
- annotation.additionalAttributes